1. Introduction
Several structural and biochemical parameters of plants, such as Gap Fraction, leaf chlorophylls a+b and carotenoids contents (
and Car), leaf mass per area (LMA), or leaf equivalent water thickness (EWT), are recognized indicators of plants’ health status [
1,
2,
3,
4]. They influence many biological and physical processes such as photosynthetic activity, nutrient cycles, gross primary production, rainfall interception, and heat fluxes [
5,
6,
7].
While field and laboratory measurements can only provide limited information on these indicators in both time and space, multispectral and hyperspectral remote sensing data have been extensively used to estimate canopy structural and biochemical parameters over large areas and can allow for recurrent measurements of the study sites [
8,
9,
10]. Hyperspectral sensors measure forest canopy reflectance using numerous spectral bands over the solar spectrum, so that slight variations of the reflected radiation can be detected. Common estimation methods of Gap Fraction and leaf biochemical properties belong to three main families: empirical-statistical methods calibrate a model to validation data acquired in-field, physical approaches rely on the inversion of radiation transfer models (RTMs) that simulate canopy reflectance, and hybrid methods bring together the fine-tuning of physically-based approaches with the flexibility of empirical-statistical methods. Baret and Buis [
11] and Verrelst et al. [
12] provide reviews of the various estimation methods, their respective difficulties, and the current solutions to try to overcome them.
In many cases, data are insufficiently available to calibrate an empirical-statistical method, and it is necessary to turn to physical or hybrid approaches and rely on RTMs. This can prove to be computationally demanding, as a consequent number of simulations could be necessary so that acceptable accuracy in variable retrieval is achieved. An important number of RTMs, either using homogeneous or heterogeneous scenes (hereafter designated as "1D" and "3D" RTMs), are available and several took part in the RAdiation transfer Model Intercomparison (RAMI) experiments [
13,
14,
15,
16]. 1D RTMs are adapted to homogeneous scenes and are by design limited in the number of possible variable parameters. While not very realistic, this makes for very short computation time and overall easier inversions for ecosystems with medium to high canopy covers. On the contrary, 3D RTMs provide a detailed description of the canopy layers and components through many variables that can be either fixed by the user (using a priori knowledge) or kept as variable parameters. This is of prime importance in particular for the modeling of sparse forests and tree–grass ecosystems that are widely distributed on Earth [
17], as the spectral contribution of the canopy to the total scene reflectance is limited and ground and shadows are more visible to the sensors. However, they most commonly rely on ray tracing methods and this added complexity can lead to a dramatic increase in computation times, which limits the sampling schemes that can realistically be considered for each variable of interest.
Unfortunately, no single best sampling scheme has been identified so far: Ali et al. [
18] considered a uniform distribution of the variables over their respective ground-truth ranges when working with INFORM [
19]; Weiss et al. [
20] drew each parameter’s values according to a distribution law which was proportional to the reflectance’s sensitivity to the parameter; Ali et al. [
21] used multivariate normal distributions and covariance matrices produced from ground truth data with INFORM; Hernández-Clemente et al. [
22] used both monovariate and multivariate random samplings with DART [
23]. Due to the computation times of 3D RTMs, testing multiple sampling schemes when building Look-Up Tables (LUTs) with tens of thousands of entries is not realistically feasible. Being able to consistently optimize the LUT sampling scheme at low time cost, either for direct LUT-based inversion or subsequent training of a machine-learning model, could therefore prove beneficial.
The aim of this study was to combine the realism of 3D RTM with the speed of 1D RTM to be able to quickly generate LUTs with several variable parameters and arbitrary sampling. To do so, the PROSAIL [
24] (1D) and DART (3D) canopy RTMs were considered. Both PROSAIL and DART were used to calibrate a model that approximates DART reflectance outputs from PROSAIL’s. The performances of this model (named PROSAIL2DART) were assessed by comparing its outputs with DART reference values. PROSAIL2DART was then used to estimate Gap Fraction, oak leaf pigment content, and oak LMA and leaf EWT over a woodland savanna. Estimations accuracies were assessed by confronting estimations with field measurements done at various stands and dates.
2. Materials and Methods
2.1. Study Site
The study site is an oak woodland savanna located in the lower foothills of the Sierra Nevada Mountains (Tonzi Ranch, latitude: 38.431°N; longitude: 120.966°W; altitude: 177 m; average slope: 1.5°). It has a Mediterranean climate alternating between mild, wet winters and hot, dry summers. Ninety percent of the overstory are Blue Oak (
Quercus douglasii—QUDO), the remaining 10% being mostly Grey Pine (
Pinus sabiniana—PISA). Blue oaks are deciduous, their leaves start to sprout in April and have been shed by November. The mean canopy cover (CC) of the site is 47%, and the mean LAI is 0.8. The understory is composed of cool season annual C3 grass species active from December to May and dry during summer and autumn. Both oak trees and grasses are active in April and May. The soil is an Auburn very rocky silt loam (Lithic haploxerepts). More detailed site information can be found in previous studies [
25,
26,
27].
As PISA only represent 10% of the overstory, the present study Gap Fraction plots contained either only QUDO or a QUDO-PISA mix with a QUDO majority (
Section 2.2.1). Leaf collection for EWT, LMA and leaf pigment content estimation were done in a pure-QUDO part of the site (
Section 2.2.2). Only QUDO were modeled in the RTM, as correctly modeling coniferous trees in 3D RTMs can be challenging [
28] and no PISA-dominant plots were included in the study (three stands are mixed and PISA canopy cover represents only 37% of the total canopy cover in the most mixed stand).
Figure 1 shows an aerial view of the site and the locations of the various plots used in this study. Picture of both QUDO and PISA as well as average dimensions of QUDO are shown in
Figure 2a,b.
2.2. Field Data
2.2.1. Gap Fraction
Field data were collected coincident with the NASA Hyperspectral Infrared Imager (HyspIRI) Mission Study Airborne Campaigns that took place in September 2013 and June 2014 and 2016 (
https://hyspiri.jpl.nasa.gov/). Several digital hemispherical photographs (DHP) were collected over multiple 60 m × 60 m plots across the study site covering all vegetation cover fractions and species composition. These plots were selected to span the full variation in species composition and canopy density. Information concerning the number of plots for each date is given in
Table 1.
From each 60 m × 60 m plot, nine DHP were taken using a Nikon Coolpix 4300 camera post sunset when no direct sunlight was visible. The DHP were taken according to the sampling patterns shown in
Figure 3. DHP were processed using CAN-EYE (
https://www6.paca.inrae.fr/can-eye). CAN-EYE calculated the Gap Fraction with azimutal and zenithal resolutions of 2.5°, using a circle of interest of 65°. The theory behind CAN-EYE estimations is described by Weiss et al. [
29] and is also available at
https://www6.paca.inrae.fr/can-eye/Documentation/Documentation. As the LAI of the site is very low, there is no significant risk of Gap Fraction saturation as this phenomenon happens for LAI higher than 5.
2.2.2. Equivalent Water Thickness, Leaf Mass Per Area, and Leaf Biochemistry Measurements
To retrieve EWT, LMA, and leaf biochemistry, a set of fully expanded leaves was collected five healthy QUDO individuals presenting a structure typical of the site. Leaves were collected from the upper, sunlit portion of the canopy. Sampling started within an hour of the timing of the overflight. Leaf samples were collected from open grown trees that were in full sunlight, as high into the canopy as possible, and from branches on the east and west sides of the tree. Attention was paid to ensure that collected leaves were healthy, and collection always occurred during dry days. Leaves were placed in a plastic bag and stored on blue ice (or in a lab refrigerator) until lab measurements could be made (<48 h). Plastic bags were weighed with a mg precision scale before going to the field. In the lab, the bags with leaves inside were weighed, with the weight difference giving leaf fresh weight. After that, the thickness of each leaf was measured using a caliper and all leaves were scanned in TIF format with 150 dpi. Leaf area was estimated using the scanned image in TOASTER software. Finally, all the leaves were put into a paper bag to dry at 65 degrees Celsius until the weight did not change when the leaves were reweighed (two to three days) to obtain leaf dry weight. Finally, EWT and LMA were calculated according to Equations (
1) and (
2).
The methodology for leaf
and Car retrieval has been described by Miraglio et al. [
30].
2.2.3. Trunk Reflectances
Tree trunk reflectances were collected from the five individuals from the leaf biochemistry collection and measured with an Analytical Spectral Device (ASD; ASD Inc., Boulder, CO, USA) contact probe. A spectralon panel was used for calibration purposes before every acquisition. Trunk reflectances were obtained over the 0.350 to 2.500 m spectral range. Small portions of the trunk were collected and situated in a horizontal surface to facilitate the measurement.
2.2.4. Airborne Hyperspectral Remote Sensing Data
AVIRIS-C hyperspectral data are processed and delivered by NASA Jet Propulsion Laboratory (JPL;
http://aviris.jpl.nasa.gov).
Table 2 gives information about the date of the acquisitions used in this study. Images were acquired at nadir 20 km above the ground within ±1 h of the solar noon to avoid spectral directional effects. Preprocessing steps provided by NASA JPL included radiometric calibration, geometrical orthorectification, nearest neighbor spatial resampling at 18 m, and atmospherical correction performed with ATREM [
31], in order to retrieve surface reflectance. The AVIRIS-C images used in this study were co-registered and spectral temporal corrections were applied using the same protocol as in Miraglio et al. [
30]. Hyperspectral images from April and October 2014 were also available but could not be used in this study, as grass was still green in April, which would have led to additional complexity when doing Gap Fraction and leaf biochemistry estimations, and a fire plume was above the site in October at the time of the airborne acquisitions.
2.3. PROSAIL2DART
2.3.1. Methodology
Both PROSAIL and DART were used to compute scene reflectances over a calibration (LAI,
, Car, LMA, EWT) grid hereafter designated as
. Band reflectance ratios
were computed for each point of the grid and used to calibrate linear 5-D interpolators. These interpolators are used to transform any reflectance obtained with PROSAIL into a reflectance similar to what DART would have obtained. For each date, one 5-D interpolator was computed for each (CC, understory reflectance, Anthocyanins content (
)) triplet as these parameters were either not possible to model in PROSAIL (CC) or to limit the uncertainties (understory reflectance,
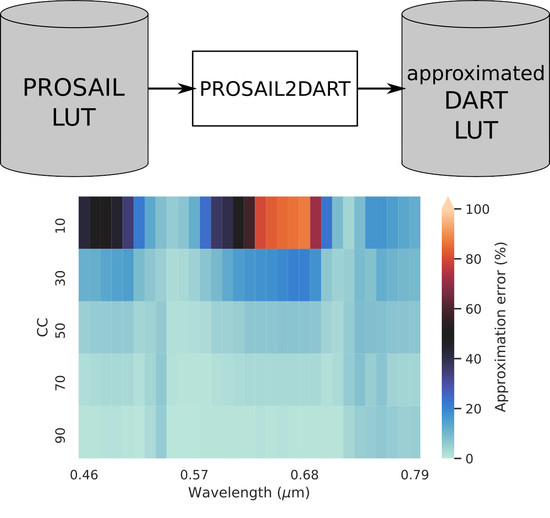
). This calibration/transformation method is hereafter called PROSAIL2DART (P2D). A diagram of the methodology is shown in
Figure 4.
2.3.2. RTM Parametrization
DART
DART version 5.7.3v1078 was used to simulate canopy reflectances. DART is a radiation transfer model able to simulate light interactions and multiple scattering effects within a 3D scene, including the topography and the atmosphere. DART includes PROSPECT to model leaf reflectance and transmittance. A precise description of the DART model can be found in Gastellu-Etchegorry et al. [
32] and Gastellu-Etchegorry et al. [
23]. Trees are defined by structural parameters such as the shape and size of their crown or the distribution and optical properties of their leaves.
The scene modeling done in this study is based on the same simplified forest representation as the one done in Miraglio et al. [
30]: canopy is represented with 4 lollipop trees and the ground is modeled as a lambertian surface, the reflectance of which was extracted from AVIRIS-C images, as this previously proved sufficient to estimate both LAI and leaf pigment content at the AVIRIS-C spatial resolution. Different soil reflectance were considered: from the sets of pure soil pixels extracted from open parts of the site, mean and mean ± standard deviation reflectances were used to build the LUTs in order to better take into account possible ground reflectance variations over the site. For simplicity purposes and to ensure that (i) the (LAI,
, Car, LMA, EWT) space that could be covered by P2D was an hyperrectangle and (ii) the density of calibration samples was uniform over this space,
followed a regular sampling scheme.
Table 3 and
Table 4 describe the various inputs used to create the DART scenes.
The Gap Fractions of the DART scenes were computed for all combinations of CC and LAI. It was retrieved using the DART 3D Radiative Budget tool, by considering the percentage of diffuse illumination intercepted by the ground. The specific DART parameters to obtain these results are
illumination using a single wavelength,
no radiative transfer in the atmosphere,
SKYL (atmospheric scattering of sun radiance) set to 1,
number of iterations set to 0, and
smaller mesh size of irradiance sources set to 0.005 m
Gap Fraction can be considered a function of CC an LAI, i.e., . Therefore, when generating the P2D, fine LUTs Gap Fractions were derived from the (CC, LAI) values by linear interpolation, using the and values as reference.
PROSAIL
The 4SAIL version of PROSAIL was used in the present study, using a Python wrapper (
https://github.com/jgomezdans/prosail, DOI: 10.5281/zenodo.2574925). PROSAIL combines the leaf model PROSPECT with the 1-D turbid canopy RTM SAIL. A thorough description and history of the PROSAIL model can be found in Berger et al. [
33]. Leaf angle distribution (LAD) is not a direct input of PROSAIL and both the average leaf slope LIDFa and the associated distribution bimodality LIDFb must be given. A spherical LAD is obtained with LIDFa =
and LIDFb =
. Ground reflectance was the same reflectance as that given to DART, as were the solar zenith and azimuth angles. LAI variations were the same as those given to DART.
PROSPECT
Leaf optical properties were simulated using the PROSPECT model, which is implemented in DART and PROSAIL. PROSPECT-D [
34] was used in this study, and a small
was introduced as a possible case for September to take into account possible leaf senescence. The leaf structure parameter N was set to 1.8. The PROSPECT specific parameters considered in this study are given in
Table 4.
2.3.3. Error Assessment
As the P2D linear interpolators were calibrated on a regular grid, the maximum differences between P2D and DART reflectances are located at the centers of the hypercubes defined by the grid. The P2D approach will be validated if the difference between P2D and DART reflectances at the hypercube center are negligible.
Let
designate the set of the hypercubes centers, which correspond to the
grid. Let
and
be the reflectances computed by P2D and DART on
points and
the reflectances computed by DART on
points, which correspond to the hypercubes corners. The P2D approximation was evaluated using the
E ratio, computed as
with
i the hypercube identifier and
j the hypercube corner identifier. This ratio was designed to compare the reflectance distance between P2D and DART at the hypercube center (maximum error) with the reflectance distance between the hypercube center and corners obtained through DART. A value close to 100 indicates that the P2D error is similar to the smallest difference between the hypercube’s center and corners, while a value close to 0 indicates that the P2D error is negligible. An illustration of the P2D validation methodology is presented
Figure 5. An
E value lower than 50 indicates that the P2D approximation is closer to the hypercube center than the corners. A condition that
should be non-negligible (>0.001 when reflectance range is 0 to 1) was used as variables do not necessarily have an influence at all wavelengths, which could lead to close to zero differences between some corners and the center and make
E diverge erroneously.
2.4. Fine Lut Building
P2D was subsequently used to generate a fine DART-like LUT for each AVIRIS-C image, following a Latin Hypercube sampling. The correlation between
and Car visible in the field data was taken into account by constraining the Car values around 2.5 times the standard deviation between field data and the regression line (see
Figure 6). For LAI,
, LMA and EWT are the boundaries of the Latin Hypercube corresponding to those of the calibration LUTs presented in
Table 4. Only CC 30 to 90% were considered, and for every ground reflectance and
value, 50,000 cases were generated and distributed equally among the CC. Therefore, June 2013, 2014, and 2016 fine LUTs each have 150,000 entries, and September 2013 LUT has 300,000 entries.
2.5. Lut-Based Inversions
To assess the performance improvement when using P2D instead of simply using DART with a regular sampling scheme, both DART
and P2D fine LUTs were used to retrieve Gap Fraction and leaf biochemistry. LUT-based approaches consist in finding the simulated reflectance
that is the most similar to the measured one,
y, according to a cost function. Several cost functions were selected for this study: root mean square error (RMSE; Equation (
4)), spectral angle mapper (SAM; Equation (
5)), and vegetation index (VI) differences (
; Equation (
6)).
RMSE and SAM were computed using variable-specific spectral intervals. The Gap Fraction interval covered the near-infrared (NIR) and short wavelength infrared (SWIR) (INT GAP, 0.8–2.45 m). Intervals for and Car were parts of the visible range (INT CAB, 0.5–0.75; INT CAR, 0.5–0.55 m), while those used for LMA and EWT were parts of the NIR and SWIR (INT LMA, 0.8–1.3; INT EWT, 1.3–2.45 m). The spectral intervals were chosen based on their sensitivity to the variables of interest according to the results of a Sobol sensitivity analysis on the DART LUTs (not shown).
Before LUT-based inversion, VI capabilities to estimate the variables of interest were assessed by fitting a function between VI and variables ( for biochemistry and for Gap Fraction, with v the variable’s value). If no relationship could be found, the VI was not considered for the inversion.
Table 5 shows all the cost functions tested for this study for each variable of interest, as well as the goodness of the fit of each VI when applicable. Estimation results were computed as the mean of multiple best solutions. The number of solutions considered for each LUT was 0.5% of the LUT size.
2.6. Validation Metrics
Gap fraction, leaf pigments content, EWT, and LMA estimates were compared with the field measurements available using the following criteria; total RMSE, systematic and unsystematic RMSE (Willmott [
53]), the model performance index
(Willmott et al. [
54]), and
of the predicted vs. measured regression line.
Concerning Gap Fraction, each validation point was the average of Gap Fraction values derived from hemispherical pictures taken within a 60 m × 60 m plot (see
Section 2.2.1). Direct comparison between pixel-estimated value and validation data can be inappropriate as the area covered by the DHP of a plot is wider than the AVIRIS-C pixel (18 m × 18 m). Therefore, validation values were compared to the average of the Gap Fractions estimated over a 3 × 3 pixels windows centered on the pixel corresponding to the plot center, similarly to the method employed in Miraglio et al. [
30].
Biochemistry validation data were obtained at the leaf scale, for one tree in each validation pixel. It was assumed that biochemistry estimations could be directly extracted from the pixels associated with the acquisition positions.
3. Results
3.1. Comparison between Aviris-C and Dart Reflectances
Figure 7 shows the validation pixels’ reflectances and compares them to the reflectance extrema found in their corresponding LUT. For June 2013, September 2013 and June 2016, all pure pixel reflectances fall within the extrema of the LUTs whatever the wavelength. One reflectance from a mixed plot is severely out of the boundaries of the LUT, while up to 8 other mixed plot reflectances are slightly below the LUT minima between 0.9 and 1.6
m, for a total of at most 9 reflectances below the LUT minima at some wavelengths out of 63 for June 2016. For June 2014, almost all reflectances including those from mixed pixels also fall within the extrema: of the 68 pixel reflectances, 6 are below the LUT minima around 1.24
m and 1.6
m, with a maximum difference of 0.02.
3.2. PROSAIL2DART Errors
Figure 8 and
Table 6 show the evolution of the
E ratio over the CC and wavelengths. In the visible, all wavelengths are well approximated by P2D for CC ≥ 30%, with the highest
E value being 21% at 0.68
m and 30% CC. While for 10% CC, the green and NIR are also well approximated (
E < 50%), this is not the case for the blue and red regions where
E can be above 50%. In the SWIR, for 10% CC
E values are considerably below 50% for
< 1.8
m. However, higher values are found at higher wavelengths and the maximum, 46%, is obtained at 1.49
m. Estimations with either DART
or P2D LUTs only used the CC ≥ 30% cases to avoid uncertainties.
3.3. PROSAIL2DART Fine Lut Generation
It took 12,666 h (total CPU time of a server equipped with Broadwell Intel® Xeon® CPU E5-2650 v4 @ 2.20 GHz) to generate the 21,840 reflectances required to build the DART LUT dedicated to September 2013 (the most extensive LUT, as two values of anthocyanins are also considered). For comparison, once P2D was calibrated (the calibration time is negligible), it took 1.5 h (total CPU time on a computer equipped with an Intel® Core™ i5-6300HQ CPU @ 2.30 GHz) to generate the 300,000 entries of the P2D fine LUT.
3.4. Estimation Performances
Table 5 shows the best
achieved by the various VI when fitted over each LUT. Gap Fraction was very well measured by its VI, and overall so were
and Car, with only MCARI2, R515_570, and CRI550 presenting no relation with the estimates’ values. Concerning LMA and EWT, no satisfying relation could be found, and only lma_D could find a slight relation with a
of 0.5. Only the inversion methods with
higher than 0.5 were considered suitable candidates for LUT-based inversions.
Concerning Gap Fraction, both LUTs perform in equivalent manner, with good performances whatever the method (maximum is 0.78 for applied on DART and 0.77 when applied on the P2D LUT). estimations show improved performances with the P2D LUTs for all cost functions except RMSE INT CAB and , which have slightly lower . offers the best performances, with = 0.77 when applied on the P2D LUTs. For Car, the best is also obtained with the P2D LUTs with , and P2D consistently improved the . Concerning LMA and EWT, the P2D fine LUTs appear to slightly improve performances for the selected cost functions (at the exception of SAM INT LMA that decreases slightly); however, remains low. Their respective best-performing cost functions are SAM INT LMA with DART and SAM INT EWT with P2D.
3.5. Estimation Plots
Figure 9 compares estimated and field values for the various estimates of this study when using the best performing methods identified in
Section 3.4. Gap Fraction estimations present both a high
and a low RMSE (0.78 and 0.1, respectively). While it appears that one of the June 2014 mixed plots (yellow) was overestimated, other mixed plots present a similar behavior as pure QUDO plots. Similar behaviors are found for
and Car: most of the points seem to follow the first bisector, and the point with the highest
and Car values is slightly underestimated. RMSE errors in both cases remain low (4.14 and 1.05
g/cm
2, respectively). No trend between estimations and field data could be found for either LMA and EWT (their respective
are 0.14 and 0.01) and RMSE of
g/cm
2 and
cm are obtained.
5. Conclusions
The results obtained in this study demonstrated the possibility to approximate with minimal error the reflectance outputs of DART with those of PROSAIL even at low (30%) CC. For higher CC, it was shown that approximation errors were negligible. The approximation model was further used to generate extensive LUTs to estimate Gap Fraction of mixed oak and pine stands as well as leaf , Car, EWT, and LMA of oak stands in a low-foliage woodland savanna. Gap Fraction and leaf pigment content estimations presented similar or improved performances when taking advantage of the proposed model instead of only relying on DART. EWT and LMA could not be retrieved using either models.
In summary, the findings show that acceptably approximating DART results from PROSAIL is possible and that the subsequent reflectances can be successfully used for estimation purposes of even very sparse oak stands, although conclusions should also applicable to other broadleaved stands due to the elementary modeling used in the 3D RTM. This is valuable, as 1D RTMs are dramatically faster than 3D RTMs. In the exploration phase, this allows for the testing of various sampling schemes at a negligible cost for either the training of machine learning methods, that require extensive training databases, or the generation of more complex LUTs. Approximated reflectances can also directly be used as is to retrieve canopy structural and biochemical parameters with acceptable accuracy.
Due to the tree distribution within the study site and the ground sampling distance of AVIRIS-C, no pine-dominant stands could be considered for Gap Fraction and leaf biochemistry estimations and this study focused mainly on pure-oak stands. Further work is necessary to extend them to coniferous trees or mixed stands. More work is also necessary to acceptably estimate EWT and LMA of tree–grass ecosystems, possibly by improving the soil realism by modeling the grass layer [
66] or the tree representation with the inclusion of detailed trunk structures [
16] within the 3D RTM.