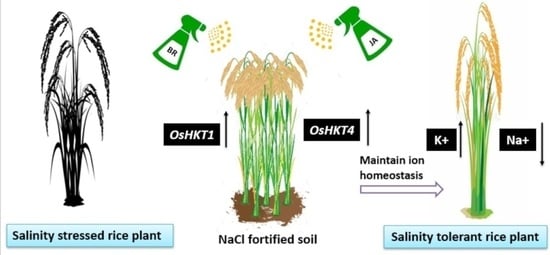
Transcriptome-Wide Analysis Revealed the Potential of the High-Affinity Potassium Transporter (HKT) Gene Family in Rice Salinity Tolerance via Ion Homeostasis
Abstract
:1. Introduction
2. Materials and Methods
2.1. HKT Genes Family Identification
2.2. Phylogenetic Tree and Digital Expression
2.3. Chromosomal Locations and Protein–Protein Interactions of OsHKT Genes
2.4. Gene Structure and Conserved Motif Analysis
2.5. Gene Ontology and Cis-Elements Analysis of HKT Family Genes
2.6. RNA Isolation and cDNA Synthesis
2.7. Expression Profiling of HKT Genes in Oryza sativa
2.8. Na+, K+, and Their Ratio in Tolerant (3Y9H) and Sensitive (JLY252) Cultivars
2.8.1. Experimental Method
2.8.2. Measurements of Na+, K+ Ratio
2.9. Statistical Analysis
3. Results
3.1. Phylogenetic Analysis of OsHKT
3.2. Gene Structure and Conserved Motif Analysis of OsHKT Genes
3.3. Gene Ontology of OsHKT Genes in Rice
3.4. Cis-Elements of OsHKT Genes in Rice
3.5. Protein–Protein Interactions of OsHKT
3.6. Tissue-Specific Expression Analysis of OsHKT Genes in Rice
3.7. Expression Analysis of OsHKT Genes in Rice Roots under Brassinosteroids Application
3.8. Expression Analysis of OsHKT Genes in Rice Root under Jasmonic Acid Application
3.9. Expression Analysis of OsHKT Genes in Rice Roots and Stems under Saline Conditions
3.10. Expression Analysis of OsHKT Genes in Salinity Resistant Cultivar Subjected to Seawater Stress
3.11. Na+, K+ Ratio in Tolerant Cultivar 3Y9H
3.12. Na+, K+ Ratio in the Sensitive Cultivar (JLY252)
4. Discussion
4.1. OsHKT Genes Are Distributed Widely in the Rice Genome
4.2. OsHKT Genes Control Plant Response to Salt Stress, Brassinosteroids, and JA Treatments
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Muthayya, S.; Sugimoto, J.D.; Montgomery, S.; Maberly, G.F. An overview of global rice production, supply, trade, and consumption. Ann. N. Y. Acad. Sci. 2014, 1324, 7–14. [Google Scholar] [CrossRef] [PubMed]
- Menguer, P.K.; Sperotto, R.A.; Ricachenevsky, F.K. A walk on the wild side: Oryza species as source for rice abiotic stress tolerance. Genet. Mol. Biol. 2017, 40, 238–252. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chang, L.; Hu, M.; Ning, J.; He, W.; Gao, J.; Ndjiondjop, M.-N.; Fu, Y.; Liu, F.; Sun, H.; Gu, P.; et al. The genetic control of glabrous glume during African rice domestication. J. Genet. Genom. 2022, 49, 427–436. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Mauleon, R.; Hu, Z.; Chebotarov, D.; Tai, S.; Wu, Z.; Li, M.; Zheng, T.; Fuentes, R.R.; Zhang, F.; et al. Genomic variation in 3010 diverse accessions of Asian cultivated rice. Nature 2018, 557, 43–49. [Google Scholar] [CrossRef]
- Ammiraju, J.S.; Luo, M.; Goicoechea, J.L.; Wang, W.; Kudrna, D.; Mueller, C.; Talag, J.; Kim, H.; Sisneros, N.B.; Blackmon, B.; et al. The Oryza bacterial artificial chromosome library resource: Construction and analysis of 12 deep-coverage large-insert BAC libraries that represent the 10 genome types of the genus Oryza. Genome Res. 2006, 16, 140–147. [Google Scholar] [CrossRef] [Green Version]
- Moradi, F.; Ismail, A.; Egdane, J.; Gregorio, G. Salinity tolerance of rice during reproductive development and association with tolerance at the seedling stage. Indian J. Plant Physiol. 2003, 8, 105–116. [Google Scholar]
- Hoang, T.M.L.; Tran, T.N.; Nguyen, T.K.T.; Williams, B.; Wurm, P.; Bellairs, S.; Mundree, S. Improvement of salinity stress tolerance in rice: Challenges and opportunities. Agronomy 2016, 6, 54. [Google Scholar] [CrossRef]
- Kumar, V.; Singh, A.; Mithra, S.A.; Krishnamurthy, S.; Parida, S.K.; Jain, S.; Tiwari, K.K.; Kumar, P.; Rao, A.R.; Sharma, S.K.; et al. Genome-wide association mapping of salinity tolerance in rice (Oryza sativa). DNA Res. 2015, 22, 133–145. [Google Scholar] [CrossRef] [Green Version]
- Jiang, Z.; Song, G.; Shan, X.; Wei, Z.; Liu, Y.; Jiang, C.; Jiang, Y.; Jin, F.; Li, Y. Association analysis and identification of ZmHKT1; 5 variation with salt-stress tolerance. Front. Plant Sci. 2018, 9, 1485. [Google Scholar] [CrossRef]
- Gadisa, N.; Wakgari, T. Nitrogen Uptake and Use Efficiency of Wheat (Triticum aestivum L.) Varieties as Influenced by Combined Application of Vermicompost and Nitrogen Fertilizer Rate. Am. J. Biosci. Bioeng. 2021, 9, 169–178. [Google Scholar] [CrossRef]
- Platten, J.D.; Egdane, J.A.; Ismail, A.M. Salinity tolerance, Na+ exclusion and allele mining of HKT1; 5 in Oryza sativa and O. glaberrima: Many sources, many genes, one mechanism? BMC Plant Biol. 2013, 13, 32. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ali, A.; Raddatz, N.; Aman, R.; Kim, S.; Park, H.C.; Jan, M.; Baek, D.; Khan, I.U.; Oh, D.-H.; Lee, S.Y.; et al. A single amino-acid substitution in the sodium transporter HKT1 associated with plant salt tolerance. Plant Physiol. 2016, 171, 2112–2126. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- An, D.; Chen, J.-G.; Gao, Y.-Q.; Li, X.; Chao, Z.-F.; Chen, Z.-R.; Li, Q.-Q.; Han, M.-L.; Wang, Y.-L.; Wang, Y.-F.; et al. AtHKT1 drives adaptation of Arabidopsis thaliana to salinity by reducing floral sodium content. PLoS Genet. 2017, 13, e1007086. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Almeida, P.M.; de Boer, G.-J.; de Boer, A.H. Assessment of natural variation in the first pore domain of the tomato HKT1; 2 transporter and characterization of mutated versions of SlHKT1; 2 expressed in Xenopus laevis oocytes and via complementation of the salt sensitive athkt1; 1 mutant. Front. Plant Sci. 2014, 5, 600. [Google Scholar] [CrossRef] [Green Version]
- Busoms, S.; Paajanen, P.; Marburger, S.; Bray, S.; Huang, X.-Y.; Poschenrieder, C.; Yant, L.; Salt, D.E. Fluctuating selection on migrant adaptive sodium transporter alleles in coastal Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2018, 115, E12443–E12452. [Google Scholar] [CrossRef] [Green Version]
- Munns, R.; James, R.A.; Xu, B.; Athman, A.; Conn, S.J.; Jordans, C.; Byrt, C.S.; Hare, R.A.; Tyerman, S.D.; Tester, M.; et al. Wheat grain yield on saline soils is improved by an ancestral Na+ transporter gene. Nat. Biotechnol. 2012, 30, 360–364. [Google Scholar] [CrossRef]
- Byrt, C.S.; Xu, B.; Krishnan, M.; Lightfoot, D.J.; Athman, A.; Jacobs, A.K.; Watson-Haigh, N.S.; Plett, D.; Munns, R.; Tester, M.; et al. The Na+ transporter, Ta hkt 1; 5-d, limits shoot Na+ accumulation in bread wheat. Plant J. 2014, 80, 516–526. [Google Scholar] [CrossRef] [Green Version]
- Thomson, M.J.; de Ocampo, M.; Egdane, J.; Rahman, M.A.; Sajise, A.G.; Adorada, D.L.; Tumimbang-Raiz, E.; Blumwald, E.; Seraj, Z.I.; Singh, R.K.; et al. Characterizing the Saltol quantitative trait locus for salinity tolerance in rice. Rice 2010, 3, 148–160. [Google Scholar] [CrossRef] [Green Version]
- Cotsaftis, O.; Plett, D.; Shirley, N.; Tester, M.; Hrmova, M. A two-staged model of Na+ exclusion in rice explained by 3D modeling of HKT transporters and alternative splicing. PLoS ONE 2012, 7, e39865. [Google Scholar]
- van Bezouw, R.F.; Janssen, E.M.; Ashrafuzzaman, M.; Ghahramanzadeh, R.; Kilian, B.; Graner, A.; Visser, R.G.; van der Linden, C.G. Shoot sodium exclusion in salt stressed barley (Hordeum vulgare L.) is determined by allele specific increased expression of HKT1; 5. J. Plant Physiol. 2019, 241, 153029. [Google Scholar] [CrossRef]
- Pulipati, S.; Somasundaram, S.; Rana, N.; Kumaresan, K.; Shafi, M.; Civáň, P.; Sellamuthu, G.; Jaganathan, D.; Ramaravi, P.V.; Punitha, S.; et al. Diversity of Sodium Transporter HKT1; 5 in Genus Oryza. Rice Sci. 2022, 29, 31–46. [Google Scholar] [CrossRef]
- Houston, R.D.; Bean, T.P.; Macqueen, D.J.; Gundappa, M.K.; Jin, Y.H.; Jenkins, T.L.; Selly, S.L.C.; Martin, S.A.; Stevens, J.R.; Santos, E.M.; et al. Harnessing genomics to fast-track genetic improvement in aquaculture. Nat. Rev. Genet. 2020, 21, 389–409. [Google Scholar] [CrossRef] [PubMed]
- Lamesch, P.; Berardini, T.Z.; Li, D.; Swarbreck, D.; Wilks, C.; Sasidharan, R.; Muller, R.; Dreher, K.; Alexander, D.L.; Garcia-Hernandez, M.; et al. The Arabidopsis Information Resource (TAIR): Improved gene annotation and new tools. Nucleic Acids Res. 2012, 40, D1202–D1210. [Google Scholar] [CrossRef] [PubMed]
- Lukaszewski, A.J.; Alberti, A.; Sharpe, A.; Kilian, A.; Stanca, A.M.; Keller, B.; Clavijo, B.J.; Friebe, B.; Gill, B.; Wulff, B.; et al. A chromosome-based draft sequence of the hexaploid bread wheat (Triticum aestivum L.) genome. Science 2014, 345, 1251788. [Google Scholar]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Borrill, P.; Ramirez-Gonzalez, R.; Uauy, C. expVIP: A customizable RNA-seq data analysis and visualization platform. Plant Physiol. 2016, 170, 2172–2186. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ma, J.; Yang, Y.; Luo, W.; Yang, C.; Ding, P.; Liu, Y.; Qiao, L.; Chang, Z.; Geng, H.; Wang, P.; et al. Genome-wide identification and analysis of the MADS-box gene family in bread wheat (Triticum aestivum L.). PLoS ONE 2017, 12, e0181443. [Google Scholar] [CrossRef] [Green Version]
- Ahmad, S.; Zhu, H.; Chen, Y.; Xi, C.; Shah, A.Z.; Ge, L. Comprehensive Bioinformatics and Expression Analysis of the TLP Gene Family Revealed Its Role in Regulating the Response of Oryza sativa to Nilaparvata lugens, Laodelphax striatellus, and Jinggangmycin. Agronomy 2022, 12, 1297. [Google Scholar] [CrossRef]
- Guo, A.-Y.; Zhu, Q.-H.; Chen, X.; Luo, J.-C. GSDS: A gene structure display server. Hereditas 2007, 29, 1023–1026. [Google Scholar] [CrossRef]
- Bailey, T.L.; Boden, M.; Buske, F.A.; Frith, M.; Grant, C.E.; Clementi, L.; Ren, J.; Li, W.W.; Noble, W.S. MEME SUITE: Tools for motif discovery and searching. Nucleic Acids Res. 2009, 37, W202–W208. [Google Scholar] [CrossRef]
- Somasundaram, S.; Véry, A.-A.; Vinekar, R.S.; Ishikawa, T.; Kumari, K.; Pulipati, S.; Kumaresan, K.; Corratgé-Faillie, C.; Sowdhamini, R.; Parida, A.; et al. Homology modeling identifies crucial amino-acid residues that confer higher Na+ transport capacity of OcHKT1; 5 from Oryza coarctata Roxb. Plant Cell Physiol. 2020, 61, 1321–1334. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Cao, X.-L.; Niu, J.-P. Effects of exogenous ascorbic acid on seed germination and seedling salt-tolerance of alfalfa. PLoS ONE 2021, 16, e0250926. [Google Scholar] [CrossRef] [PubMed]
- Bartha, C.; Fodorpataki, L.; Martinez-Ballesta, M.d.C.; Popescu, O.; Carvajal, M. Sodium accumulation contributes to salt stress tolerance in lettuce cultivars. J. Appl. Bot. Food Qual. 2015, 88, 42–48. [Google Scholar]
- Khan, N.; Choi, J.Y.; Nho, E.Y.; Jamila, N.; Habte, G.; Hong, J.H.; Hwang, I.M.; Kim, K.S. Determination of minor and trace elements in aromatic spices by micro-wave assisted digestion and inductively coupled plasma-mass spectrometry. Food Chem. 2014, 158, 200–206. [Google Scholar] [CrossRef]
- Letunic, I.; Bork, P. Interactive Tree Of Life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 2021, 49, W293–W296. [Google Scholar] [CrossRef]
- Sato, Y.; Takehisa, H.; Kamatsuki, K.; Minami, H.; Namiki, N.; Ikawa, H.; Ohyanagi, H.; Sugimoto, K.; Antonio, B.A.; Nagamura, Y. RiceXPro version 3.0: Expanding the informatics resource for rice transcriptome. Nucleic Acids Res. 2013, 41, D1206–D1213. [Google Scholar] [CrossRef] [Green Version]
- Rus, A.; Yokoi, S.; Sharkhuu, A.; Reddy, M.; Lee, B.-H.; Matsumoto, T.K.; Koiwa, H.; Zhu, J.-K.; Bressan, R.A.; Hasegawa, P.M. AtHKT1 is a salt tolerance determinant that controls Na+ entry into plant roots. Proc. Natl. Acad. Sci. USA 2001, 98, 14150–14155. [Google Scholar] [CrossRef] [Green Version]
- Wang, T.-B.; Gassmann, W.; Rubio, F.; Schroeder, J.I.; Glass, A.D. Rapid up-regulation of HKT1, a high-affinity potassium transporter gene, in roots of barley and wheat following withdrawal of potassium. Plant Physiol. 1998, 118, 651–659. [Google Scholar] [CrossRef] [Green Version]
- Wang, M.; Vannozzi, A.; Wang, G.; Liang, Y.-H.; Tornielli, G.B.; Zenoni, S.; Cavallini, E.; Pezzotti, M.; Cheng, Z.-M. Genome and transcriptome analysis of the grapevine (Vitis vinifera L.) WRKY gene family. Hortic. Res. 2014, 1, 14016. [Google Scholar] [CrossRef] [Green Version]
- Ren, Z.; Liu, Y.; Kang, D.; Fan, K.; Wang, C.; Wang, G.; Liu, Y. Two alternative splicing variants of maize HKT1; 1 confer salt tolerance in transgenic tobacco plants. Plant Cell Tissue Organ Cult. (PCTOC) 2015, 123, 569–578. [Google Scholar] [CrossRef]
- Chen, W.; Provart, N.J.; Glazebrook, J.; Katagiri, F.; Chang, H.-S.; Eulgem, T.; Mauch, F.; Luan, S.; Zou, G.; Whitham, S.A.; et al. Expression profile matrix of Arabidopsis transcription factor genes suggests their putative functions in response to environmental stresses. Plant Cell 2002, 14, 559–574. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Xu, G.; Yang, C.; Yang, L.; Liang, Z. Genome-wide identification and expression analysis of HKT transcription factor under salt stress in nine plant species. Ecotoxicol. Environ. Saf. 2019, 171, 435–442. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Xie, P.; Lascoux, M.; Meagher, T.R.; Liu, J. Rapidly evolving genes and stress adaptation of two desert poplars, Populus euphratica and P. pruinosa. PLoS ONE 2013, 8, e66370. [Google Scholar] [CrossRef] [Green Version]
- Azeem, F.; Zameer, R.; Rehman Rashid, M.A.; Rasul, I.; Ul-Allah, S.; Siddique, M.H.; Fiaz, S.; Raza, A.; Younas, A.; Rasool, A.; et al. Genome-wide analysis of potassium transport genes in Gossypium Raimondi suggest a role of GrHAK/KUP/KT8, GrAKT2.1 and GrAKT1.1 in response to abiotic stress. Plant Physiol. Biochem. 2022, 170, 110–122. [Google Scholar] [CrossRef]
- Riedelsberger, J.; Miller, J.K.; Valdebenito-Maturana, B.; Piñeros, M.A.; González, W.; Dreyer, I. Plant HKT Channels: An Updated View on Structure, Function and Gene Regulation. Int. J. Mol. Sci. 2021, 22, 1892. [Google Scholar] [CrossRef]
- Scherf, M.; Klingenhoff, A.; Werner, T. Highly specific localization of promoter regions in large genomic sequences by PromoterInspector: A novel context analysis approach. J. Mol. Biol. 2000, 297, 599–606. [Google Scholar] [CrossRef] [Green Version]
- Durrant, W.E.; Rowland, O.; Piedras, P.; Hammond-Kosack, K.E.; Jones, J.D. cDNA-AFLP reveals a striking overlap in race-specific resistance and wound response gene expression profiles. Plant Cell 2000, 12, 963–977. [Google Scholar] [CrossRef]
- Singh, K.B.; Foley, R.C.; Oñate-Sánchez, L. Transcription factors in plant defense and stress responses. Curr. Opin. Plant Biol. 2002, 5, 430–436. [Google Scholar] [CrossRef]
- Ren, Z.-H.; Gao, J.-P.; Li, L.-G.; Cai, X.-L.; Huang, W.; Chao, D.-Y.; Zhu, M.-Z.; Wang, Z.-Y.; Luan, S.; Lin, H.-X. A rice quantitative trait locus for salt tolerance encodes a sodium transporter. Nat. Genet. 2005, 37, 1141–1146. [Google Scholar] [CrossRef]
- Horie, T.; Yoshida, K.; Nakayama, H.; Yamada, K.; Oiki, S.; Shinmyo, A. Two types of HKT transporters with different properties of Na+ and K+ transport in Oryza sativa. Plant J. 2001, 27, 129–138. [Google Scholar] [CrossRef]
- Lin, H.; Zhu, M.; Yano, M.; Gao, J.; Liang, Z.; Su, W.; Hu, X.; Ren, Z.; Chao, D.Y. QTLs for Na+ and K+ uptake of the shoots and roots controlling rice salt tolerance. Theor. Appl. Genet. 2004, 108, 253–260. [Google Scholar] [CrossRef] [PubMed]
- Imran, S.; Horie, T.; Katsuhara, M. Expression and Ion Transport Activity of Rice OsHKT1;1 Variants. Plants 2019, 9, 16. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, J.; Nan, N.; Li, N.; Liu, Y.; Wang, T.J.; Hwang, I.; Liu, B.; Xu, Z.Y. A DNA Methylation Reader-Chaperone Regulator-Transcription Factor Complex Activates OsHKT1;5 Expression during Salinity Stress. Plant Cell 2020, 32, 3535–3558. [Google Scholar] [CrossRef]
- Chen, N.; Tong, S.; Tang, H.; Zhang, Z.; Liu, B.; Lou, S.; Liu, J.; Liu, H.; Ma, T.; Jiang, Y. The PalERF109 transcription factor positively regulates salt tolerance via PalHKT1;2 in Populus alba var. pyramidalis. Tree Physiol. 2020, 40, 717–730. [Google Scholar] [CrossRef] [PubMed]
- Raza, A.; Tabassum, J.; Fakhar, A.Z.; Sharif, R.; Chen, H.; Zhang, C.; Ju, L.; Fotopoulos, V.; Siddique, K.H.M.; Singh, R.K.; et al. Smart reprograming of plants against salinity stress using modern biotechnological tools. Crit. Rev. Biotechnol. 2022, 15, 1–28. [Google Scholar] [CrossRef]
- Wang, R.; Jing, W.; Xiao, L.; Jin, Y.; Shen, L.; Zhang, W. The Rice High-Affinity Potassium Transporter1;1 Is Involved in Salt Tolerance and Regulated by an MYB-Type Transcription Factor. Plant Physiol. 2015, 168, 1076–1090. [Google Scholar] [CrossRef] [Green Version]
- Wu, H.; Shabala, L.; Zhou, M.; Su, N.; Wu, Q.; Ul-Haq, T.; Zhu, J.; Mancuso, S.; Azzarello, E.; Shabala, S. Root vacuolar Na+ sequestration but not exclusion from uptake correlates with barley salt tolerance. Plant J. 2019, 100, 55–67. [Google Scholar] [CrossRef]
- Zhang, H.; Feng, H.; Zhang, J.; Ge, R.; Zhang, L.; Wang, Y.; Li, L.; Wei, J.; Li, R. Emerging crosstalk between two signaling pathways coordinates K+ and Na+ homeostasis in the halophyte Hordeum brevisubulatum. J. Exp. Bot. 2020, 71, 4345–4358. [Google Scholar] [CrossRef]
- Ma, Q.; Hu, J.; Zhou, X.R.; Yuan, H.J.; Kumar, T.; Luan, S.; Wang, S.M. ZxAKT1 is essential for K+ uptake and K+/Na+ homeostasis in the succulent xerophyte Zygophyllum xanthoxylum. Plant J. 2017, 90, 48–60. [Google Scholar] [CrossRef] [Green Version]













| Gene Name | Locus ID | CDS | Chr. Position | AA | MW | PI | SL |
|---|---|---|---|---|---|---|---|
| OsHKT1 | LOC_Os06g48,810 | 1593 | Chr6- 29,541,219-29,538,934 | 531 | 59,295.0781 | 9.82610035 | PM |
| OsHKT3 | LOC_Os01g34,850 | 1530 | Chr1- 19,242,042-19,243,853 | 510 | 56,374.3789 | 9.11009979 | PM |
| OsHKT4 | LOC_Os04g51,820 | 1659 | Chr4- 30,727,084-30,724,244 | 553 | 61,862.4492 | 8.86629963 | PM |
| OsHKT6 | LOC_Os02g07,830 | 1596 | Chr2- 4,103,333-4,105,657 | 532 | 59,304.5312 | 9.78820038 | PM |
| OsHKT7 | LOC_Os04g51,830 | 1503 | Chr4- 30,739,334-30,734,183 | 501 | 54,239.3008 | 8.87370014 | PM |
| OsHKT8 | LOC_Os01g20,160 | 1665 | Chr1- 11,463,442-11,458,955 | 555 | 60,218.2812 | 8.68159962 | PM |
| OsHKT9 | LOC_Os06g48,800 | 1530 | Chr6- 29,536,553-29,534,805 | 510 | 56,116.7188 | 8.54440022 | PM |
| Reference Protein | Predicted Interactive Partner | Annotation and Putative Function |
|---|---|---|
| OsHKT1 | NHX2 | Putative Na+/H+ antiporter; Sodium/hydrogen exchanger. |
| NHX1 | Sodium/hydrogen exchanger; Belongs to the monovalent cation: proton antiporter 1 (CPA1) transporter (TC 2.A.36) family. | |
| P5CS1 | Delta-1-pyrroline-5-carboxylate synthase Glutamate 5-kinase Gamma-glutamyl phosphate reductase; P5CS plays a key role in proline biosynthesis, leading to osmoregulation in plants. Involved in abiotic stress tolerance. | |
| HAK1 | Potassium transporter 1; High-affinity potassium transporter. Additionally, transport rubidium, with the same affinity and cesium, with a lower affinity; which belongs to the HAK/KUP transporter (TC 2.A.72.3) family. | |
| OsJ_04382 | Putative BTB and TAZ domain protein. | |
| TPKC | Two pore potassium channel c; Inward-rectifying potassium channel; Belongs to the two-pore domain potassium channel (TC 1.A.1.7) family. | |
| OsJ_30786 | Hsp20/alpha crystalline family protein expressed; Belongs to the small heat shock protein (HSP20) family. |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hussain, S.; Zhang, R.; Liu, S.; Li, R.; Zhou, Y.; Chen, Y.; Hou, H.; Dai, Q. Transcriptome-Wide Analysis Revealed the Potential of the High-Affinity Potassium Transporter (HKT) Gene Family in Rice Salinity Tolerance via Ion Homeostasis. Bioengineering 2022, 9, 410. https://doi.org/10.3390/bioengineering9090410
Hussain S, Zhang R, Liu S, Li R, Zhou Y, Chen Y, Hou H, Dai Q. Transcriptome-Wide Analysis Revealed the Potential of the High-Affinity Potassium Transporter (HKT) Gene Family in Rice Salinity Tolerance via Ion Homeostasis. Bioengineering. 2022; 9(9):410. https://doi.org/10.3390/bioengineering9090410
Chicago/Turabian StyleHussain, Shahid, Rui Zhang, Shuli Liu, Rongkai Li, Yicheng Zhou, Yinglong Chen, Hongyan Hou, and Qigen Dai. 2022. "Transcriptome-Wide Analysis Revealed the Potential of the High-Affinity Potassium Transporter (HKT) Gene Family in Rice Salinity Tolerance via Ion Homeostasis" Bioengineering 9, no. 9: 410. https://doi.org/10.3390/bioengineering9090410