Plastome Characterization and Phylogenomic Analysis Yield New Insights into the Evolutionary Relationships among the Species of the Subgenus Bryocles (Hosta; Asparagaceae) in East Asia
Abstract
:1. Introduction
2. Results
2.1. Genome Size and Features
2.2. Comparative Plastome Analyses
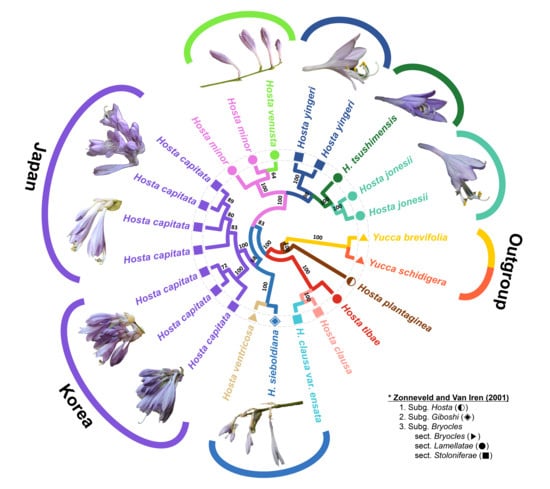
2.3. Plastome Phylogenomic Analysis of Hosta in East Asia
3. Discussion
3.1. Plastome Evolution in Subgenus Bryocles
3.2. Phylogenetic Relationships among Species of the Bryocles Subgenus in East Asia
4. Materials and Methods
4.1. Plant Materials
4.2. DNA Extraction, Sequencing, Plastome Assembly, and Annotation
4.3. Comparative Plastome Analysis
4.4. Phylogenetic Analysis
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Fujita, N. The genus Hosta (Liliaceae) in Japan. Acta Phytotax. Geobot. 1976, 27, 66–96. [Google Scholar]
- Schmid, W.G. The Genus Hosta-Giboshi Zoku; Batsford/Timber Press: London, UK, 1991. [Google Scholar]
- Chen, X.; Boufford, D.E. Hosta . In Flora of China; Wu, Z.Y., Raven, P.H., Eds.; Science Press: Beijing, China, 2000; Volume 24, pp. 204–205. [Google Scholar]
- Zonneveld, B.J.M.; Van Iren, F. Genome size and pollen viability as taxonomic criteria: Application to the genus Hosta. Plant Biol. 2001, 3, 176–185. [Google Scholar] [CrossRef]
- Chung, Y.C. Hosta . In The Genera of Vascular Plants of Korea; Park, C.W., Ed.; Academy Publishing Co.: Seoul, Korea, 2007; pp. 1304–1306. [Google Scholar]
- Tamura, M.N.; Fujita, N. Hosta . In Flora of Japan; Iwatsuki, K., Boufford, D.E., Ohba, H., Eds.; Kodansha Ltd.: Tokyo, Japan, 2016; Volume IV, pp. 140–147. [Google Scholar]
- The American Hosta Society. Genus Hosta Registration Publications. The American Hosta Society. 2020. Available online: https://www.hostaregistrar.org/hosta_registration_lists.html (accessed on 20 July 2021).
- Mimaki, Y.; Kameyama, A.; Kuroda, M.; Sashida, Y.; Hirano, T.; Oka, K.; Koike, K.; Nikaido, T. Steroidal glycosides from the underground parts of Hosta plantaginea var. japonica and their cytostatic activity on leukaemia HL-60 cells. Phytochemistry 1997, 44, 305–310. [Google Scholar] [CrossRef]
- Li, R.; Wang, M.-Y.; Li, X.-B. Chemical constituents and biological activities of genus Hosta (Liliaceae). J. Med. Plant Res. 2012, 6, 2704–2713. [Google Scholar] [CrossRef] [Green Version]
- Yang, L.; Jiang, S.T.; Zhou, Q.G.; Zhong, G.Y.; He, J.W. Chemical constituents from the flower of Hosta plantaginea with cyclooxygenases inhibition and antioxidant activities and their chemotaxonomic significance. Molecules 2017, 22, 1825. [Google Scholar] [CrossRef] [Green Version]
- Chu, H.B.; Li, N.N.; Zhang, Z.P.; Hu, X.Y.; Yu, C.Y.; Hua, L. Steroidal glycosides from the underground parts of Hosta ventricosa and their anti-inflammatory activities in mice. Nat. Prod. Res. 2019, 35, 1766–1774. [Google Scholar] [CrossRef] [PubMed]
- Jo, H.; Kim, M. A taxonomic study of the genus Hosta in Korea. Korean J. Plant Taxon. 2017, 47, 27–45. [Google Scholar] [CrossRef] [Green Version]
- Maekawa, F. The genus Hosta. J. Fac. Sci. Imp. Univ. Tokyo Sect. 3 Bot. 1940, 5, 317–425. [Google Scholar]
- Chung, Y.C.; Chung, Y.H. A taxonomic study of the genus Hosta in Korea. Korean J. Plant Taxon. 1988, 18, 161–172. [Google Scholar] [CrossRef]
- Chung, M.G. A Biosystemaitc Study on the Genus Hosta (Liliaceae/Funkiaceae) in Korea and Tsushima Island of Japan. Ph.D. Thesis, University of Georgia, Athens, GA, USA, 1990. [Google Scholar]
- Chung, M.G. Genetic structure in Korea populations of Hosta capitata (Liliaceae). J. Plant Biol. 1994, 7, 277–284. [Google Scholar]
- Chung, M.G. Low levels of genetic diversity within populations of Hosta clausa (Liliaceae). Plant Species Biol. 1994, 9, 177–182. [Google Scholar] [CrossRef]
- Chung, M.G. Genetic variation and population structure in Korean endemic species: III. Hosta minor (Liliaceae). J. Plant Res. 1994, 107, 377–383. [Google Scholar] [CrossRef]
- Chung, M.G. Genetic diversity in two island endemics, Hosta venusta and H. tsushimensis (Liliaceae). J. Jpn. Bot. 1995, 70, 322–327. [Google Scholar]
- Chung, M.G.; Kim, J.W. The genus Hosta Tratt. (Liliaceae) in Korea. SIDA 1991, 14, 411–420. [Google Scholar]
- Chung, M.G.; Hamrick, J.L.; Jones, S.B.; Derda, G.S. Isozyme variation within and among populations of Hosta (Liliaceae) in Korea. Syst. Bot. 1991, 16, 667–684. [Google Scholar] [CrossRef]
- Chung, M.G.; Jones, S.B.; Hamrick, J.L.; Chung, G.G. Morphometric and isozyme analysis of the genus Hosta (Liliaceae) in Korea. Plant Species Biol. 1991, 6, 55–69. [Google Scholar] [CrossRef]
- Lee, S.R.; Kim, K.; Lee, B.Y.; Lim, C.E. Complete chloroplast genomes of all six Hosta species occurring in Korea: Molecular structure, comparative, and phylogenetic analyses. BMC Genom. 2019, 20, 833. [Google Scholar] [CrossRef]
- Yoo, M.-J.; Lee, B.-Y.; Kim, S.; Lim, C.E. Phylogenomics with Hyb-Seq unravels Korean Hosta evolution. Front. Plant Sci. 2021, 12, 645735. [Google Scholar] [CrossRef]
- Li, P.; Lu, R.-S.; Xu, W.-Q.; Ohi-Toma, T.; Cai, M.-Q.; Qiu, Y.-X.; Cameron, K.M.; Fu, C.-X. Comparative genomics and phylogenomics of East Asian tulips (Amana, Liliaceae). Front. Plant Sci. 2017, 8, 451. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jeon, J.-H.; Kim, S.-C. Comparative analysis of the complete chloroplast genome sequences of three closely related east-Asian wild roses (Rosa sect. Synstylae; Rosaceae). Genes 2019, 10, 23. [Google Scholar] [CrossRef] [Green Version]
- Yang, J.Y.; Takayama, K.; Pak, J.-H.; Kim, S.-C. Comparison of the whole-plastome sequence between the Bonin Islands endemic Rubus boninensis and its close relative, Rubus trifidus (Rosaceae), in the southern Korean Peninsula. Genes 2019, 10, 774. [Google Scholar] [CrossRef] [Green Version]
- Yang, J.Y.; Takayama, K.; Youn, J.-S.; Pak, J.-H.; Kim, S.-C. Plastome characterization and phylogenomics of East Asian beeches with a special emphasis on Fagus multinervis on Ulleung Island, Korea. Genes 2020, 11, 1338. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.Y.; Kang, G.-H.; Pak, J.-H.; Kim, S.-C. Characterization and comparison of two complete plastomes of Rosaceae species (Potentilla dickinsii var. glabrata and Spiraea insularis) endemic to Ulleung Island, Korea. Int. J. Mol. Sci. 2020, 21, 4933. [Google Scholar] [CrossRef]
- Kim, S.-H.; Yang, J.Y.; Park, J.; Yamada, T.; Maki, M.; Kim, S.-C. Comparison of whole plastome sequences between thermogenic skunk cabbage Symplocarpus renifolius and nonthermogenic S. nipponicus (Orontioideae; Araceae) in East Asia. Int. J. Mol. Sci. 2019, 20, 4678. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kim, G.-B.; Lim, C.E.; Kim, J.-S.; Kim, K.; Lee, J.H.; Yu, H.-J.; Mun, J.-H. Comparative chloroplast genome analysis of Artemisia (Asteraceae) in East Asia: Insights into evolutionary divergence and phylogenomic implications. BMC Genom. 2020, 21, 415. [Google Scholar] [CrossRef]
- Floden, A.; Schilling, E.E. Using phylogenomics to reconstruct phylogenetic relationships within tribe Polygonateae (Asparagaceae), with a special focus on Polygonatum. Mol. Phylogenet. Evol. 2018, 129, 202–213. [Google Scholar] [CrossRef]
- Gao, F.; Chen, C.; Arab, D.A.; Du, Z.; He, Y.; Ho, S.Y.W. EasyCodeML: A visual tool for analysis of selection using CodeML. Ecol. Evol. 2019, 9, 3891–3898. [Google Scholar] [CrossRef] [Green Version]
- Yang, Z. PAML: A program package for phylogenetic analysis by maximum likelihood. Bioinformatics 1997, 13, 555–556. [Google Scholar] [CrossRef]
- Kim, S.-H.; Cho, C.H.; Yoon, H.S.; Jeon, J.; Kim, S.-C. Characterization of the complete chloroplast genome of Forsythia saxatilis (Oleaceae), a vulnerable calcicolus species endemic to Korea. Conserv. Genet. Resour. 2018, 10, 723–726. [Google Scholar] [CrossRef]
- Yang, J.Y.; Pak, J.-H.; Kim, S.-C. The plastome sequence of Ulleung Island endemic Rubus takesimensis in Korea: Insight into chloroplast genome evolution in genus Rubus (Rosaceae). Gene 2018, 668, 221–229. [Google Scholar] [CrossRef] [PubMed]
- Cho, M.-S.; Kim, J.H.; Yamada, T.; Maki, M.; Kim, S.-C. Plastome characterization and comparative analyses of wild crabapples (Malus baccata and M. toringo): Insights into infraspecific plastome variation and phylogenetic relationships. Tree Genet. Genomes 2021, in press. [Google Scholar] [CrossRef]
- Morton, B.R. Selection on the codon bias of chloroplast and cyanelle genes in different plant and algal lineages. J. Mol. Evol. 1998, 46, 449–459. [Google Scholar] [CrossRef]
- Gu, W.; Zhou, T.; Ma, J.; Sun, X.; Lu, Z. The relationship between synonymous codon usage and protein structure in Escherichia coli and Homo sapiens. BioSystems 2004, 73, 89–97. [Google Scholar] [CrossRef] [PubMed]
- Nie, X.; Deng, P.; Liu, P.; Du, X.; You, F.M.; Song, W. Comparative analysis of codon usage patterns in chloroplast genomes of the Asteraceae family. Plant Mol. Biol. Rep. 2014, 32, 828–840. [Google Scholar] [CrossRef]
- Ravi, V.; Khurana, J.P.; Tyagi, A.K.; Khurana, P. An update on chloroplast genomes. Plant Syst. Evol. 2008, 270, 101–122. [Google Scholar] [CrossRef]
- Rabah, S.O.; Lee, C.; Hajrah, N.H.; Makki, R.M.; Alharby, H.F.; Alhebshi, A.M.; Sabir, J.S.M.; Jansen, R.K.; Ruhlman, T.A. Plastome sequencing of ten nonmodel crop species uncovers a large insertion of mitochondrial DNA in cashew. Plant Genome 2017, 10, 3. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pinard, D.; Myburg, A.A.; Mizrachi, E. The plastid and mitochondrial genomes of Eucalyptus grandis. BMC Genom. 2019, 20, 132. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.Y.; Chiang, Y.-C.; Hsu, T.-W.; Pak, J.-H.; Kim, S.-C. Characterization and comparative analysis among plastome sequences of eight endemic Rubus (Rosaceae) species in Taiwan. Sci. Rep. 2021, 11, 1152. [Google Scholar] [CrossRef] [PubMed]
- Curtis, S.E.; Clegg, M.T. Molecular evolution of chloroplast DNA sequences. Mol. Biol. Evol. 1984, 1, 291–301. [Google Scholar] [PubMed] [Green Version]
- Palmer, J.D. Comparative organization of chloroplast genomes. Annu. Rev. Genet. 1985, 19, 325–354. [Google Scholar] [CrossRef]
- Redwan, R.M.; Saidin, A.; Kumar, S.V. Complete chloroplast genome sequence of MD-2 pineapple and its comparative analysis among nine other plants from the subclass Commelinidae. BMC Plant Biol. 2015, 15, 196. [Google Scholar] [CrossRef] [Green Version]
- Sloan, D.B.; Triant, D.A.; Forrester, N.J.; Bergner, L.M.; Wu, M.; Taylor, D.R. A recurring syndrome of accelerated plastid genome evolution in the angiosperm tribe Sileneae (Caryophyllaceae). Mol. Phylogenet. Evol. 2014, 72, 82–89. [Google Scholar] [CrossRef]
- Cheng, H.; Li, J.; Zhang, H.; Cai, B.; Gao, Z.; Qiao, Y.; Mi, L. The complete chloroplast genome sequence of strawberry (Fragaria × ananassa Duch.) and comparison with related species of Rosaceae. PeerJ 2017, 5, e3919. [Google Scholar] [CrossRef] [Green Version]
- Jiang, P.; Shi, F.X.; Li, M.R.; Liu, B.; Wen, J.; Xiao, H.X.; Li, L.F. Positive selection driving cytoplasmic genome evolution of the medicinally important ginseng plant genus Panax. Front. Plant Sci. 2018, 9, 359. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Lou, G.; Cai, X.; Zhang, B.; Cheng, Y.; Wang, H. Comparison of the complete plastomes and the phylogenetic analysis of Paulownia species. Sci. Rep. 2020, 10, 2225. [Google Scholar] [CrossRef] [Green Version]
- Piot, A.; Hackel, J.; Christin, P.A.; Besnard, G. One-third of the plastid genes evolved under positive selection in PACMAD grasses. Planta 2018, 247, 255–266. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Zhang, H.; Jiang, M.; Chen, H.; Huang, L.; Liu, C. Complete plastome sequence of Iodes cirrhosa Turcz., the first in the Icacinaceae, comparative genomic analyses and possible split of Idoes species in response to climate changes. PeerJ 2019, 7, e6663. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Liu, F.; Yang, D.-G.; Li, W.; Zhou, X.-J.; Pei, X.-Y.; Liu, Y.-G.; He, K.-L.; Zhang, W.-S.; Ren, Z.-Y.; et al. Comparative chloroplast genomics of Gossypium species: Insights into repeat sequence variations and phylogeny. Front. Plant Sci. 2018, 9, 376. [Google Scholar] [CrossRef] [Green Version]
- Burri, R.; Salamin, N.; Studer, R.A.; Roulin, A.; Fumagalli, L. Adaptive divergence of ancient gene duplicates in the avian MHC class II beta. Mol. Biol. Evol. 2010, 27, 2360–2374. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sauve, R.J.; Zhou, S.P.; Yu, Y.C.; Schmid, W.G. Randomly amplified polymorphic DNA analysis in the genus Hosta. HortScience 2005, 40, 1243–1245. [Google Scholar] [CrossRef] [Green Version]
- Ohba, H.; Ishii, N.; Saito, S. Biogeography of Tsushima Island. In Report on the Natural Resource Investigation of Tsushima Island; Nature of Tsushima Island: Nagasaki, Japan, 1987; pp. 259–271. [Google Scholar]
- Fujita, N.; Tamura, N.M. Two new varieties and one change of status in Hosta (Asparagaceae). Acta Phytotaxon. Geobot. 2008, 59, 31–36. [Google Scholar]
- Chung, M.G.; Jones, S.B. Pollen morphology of Hosta Tratt. (Funkiaceae) and related genera. Bull. Torrey Bot. Club 1989, 116, 31–44. [Google Scholar] [CrossRef]
- Kaneko, K. Cytological studies on some species of Hosta. III. Karyotypes of H. clausa, H. clausa var. normalis and H. ventricosa. Bot. Mag. 1968, 81, 396–403. [Google Scholar] [CrossRef] [Green Version]
- Rieseberg, L.H.; Brouillet, L. Are many plant species paraphyletic? Taxon 1994, 43, 21–32. [Google Scholar] [CrossRef]
- Zerbino, D.R.; Birney, E. Velvet: Algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 2008, 8, 821–829. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lowe, T.M.; Eddy, S.R. tRNAscan-SE: A program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 1997, 25, 955–964. [Google Scholar] [CrossRef] [PubMed]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C.; et al. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef]
- Greiner, S.; Lehwark, P.; Bock, R. Organellar Genome DRAW (OGDRAW) version 1.3.1: Expanded toolkit for the graphical visualization of organellar genomes. Nucleic Acids Res. 2019, 47, W59–W64. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brudno, M.; Malde, S.; Poliakov, A.; Do, C.B.; Couronne, O.; Dubchak, I.; Batzoglou, S. Global alignment: Finding rearrangements during alignment. Bioinformatics 2003, 19, i54–i62. [Google Scholar] [CrossRef] [Green Version]
- Frazer, K.A.; Pachter, L.; Poliakov, A.; Rubin, E.M.; Dubchak, I. VISTA: Computational tools for comparative genomics. Nucleic Acids Res. 2004, 32, W273–W279. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software v7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [Green Version]
- Rozas, J.; Ferrer-Mata, A.; Sánchez-Del Barrio, J.C.; Guirao-Rico, S.; Librado, P.; Ramos-Onsins, S.E.; Sánchez-Garcia, A. DnaSP v6: DNA sequence polymorphism analysis of large datasets. Mol. Biol. Evol. 2017, 34, 3299–3302. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [Green Version]
- Sharp, P.M.; Li, W.H. The codon adaptation index-A measure of directional synonymous codon usage bias, and its potential applications. Nucleic Acids Res. 1987, 15, 1281–1295. [Google Scholar] [CrossRef] [Green Version]
- Kozak, M. Comparison of initiation of protein synthesis in procaryotes, eucaryotes, and organelles. Microbiol. Rev. 1983, 47, 1–45. [Google Scholar] [CrossRef] [PubMed]
- Mower, J.P. The PREP suite: Predictive RNA editors for plant mitochondrial genes, chloroplast genes and user-defined alignments. Nucleic Acids Res. 2009, 37, W253–W259. [Google Scholar] [CrossRef]
- Kimura, M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 1980, 16, 111–120. [Google Scholar] [CrossRef] [PubMed]
- Swofford, D.L. PAUP * 4.0a. Phylogenetic Analysis Using Parsimony (* and Other Methods); Sinauer Associates: Sunderland, MA, USA, 2003. [Google Scholar]
- Nguyen, L.-T.; Schmidt, H.A.; von Haeseler, A.; Minh, B.Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef]





| Taxa | Hosta capitata (Mt. BaekUn) | Hosta capitata (Mt. DaeDuck) | Hosta capitata (Iya Valley) | Hosta capitata (Mt. Rokko) | Hosta capitata (Pref. Kochi) | Hosta capitata (Pref. Miyazaki) | Hosta jonesii | Hosta sieboldiana | Hosta tibae | Hosta tsushimensis | Hosta yingeri |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Total cpDNA size (bp) | 156,419 | 156,423 | 156,682 | 156,678 | 156,675 | 156,685 | 156,724 | 156,576 | 156,626 | 156,714 | 156,755 |
| GC content (%) | 37.82 | 37.82 | 37.81 | 37.81 | 37.81 | 37.81 | 37.81 | 37.81 | 37.82 | 37.8 | 37.8 |
| LSC size (bp) /GC content (%) | 84,785 /35.89 | 84,795 /35.88 | 85,054 /35.87 | 85,054 /35.87 | 85,049 /35.87 | 85,054 /35.87 | 85,104 /35.88 | 84,952 /35.87 | 85,014 /35.9 | 85,100 /35.86 | 85,115 /35.88 |
| IR size (bp) /GC content (%) | 26,711 /42.91 | 26,711 /42.91 | 26,711 /42.91 | 26,711 /42.91 | 26,711 /42.91 | 26,711 /42.91 | 26,698 /42.93 | 26,696 /42.93 | 26,697 /42.92 | 26,698 /42.92 | 26,704 /42.91 |
| SSC size (bp) /GC content (%) | 18,205 /31.9 | 18,206 /31.9 | 18,206 /31.91 | 18,202 /31.91 | 18,204 /31.91 | 18,209 /31.89 | 18,224 /31.82 | 18,232 /31.85 | 18,218 /31.87 | 18,218 /31.87 | 18,232 /31.86 |
| Accession number | MZ919305 | MZ919306 | MZ919307 | MZ919308 | MZ919309 | MZ919310 | MZ919311 | MZ919312 | MZ919313 | MZ919314 | MZ919315 |
| Gene Name | Models | np | ln L | Likelihood Ratio Test p-Value | Positively Selected Sites |
|---|---|---|---|---|---|
| rbcL | M8 | 27 | −2050.484207 | <0.000000001 | 477 Q 0.973 * |
| M7 | 25 | −2092.011742 | |||
| rpoB | M8 | 27 | −4391.457170 | <0.000000001 | 514 N 0.962 * |
| M7 | 25 | −4436.574724 | |||
| rpoC2 | M8 | 27 | −5749.647291 | <0.000000001 | 754 W 0.989 * |
| M7 | 25 | −5784.027018 | |||
| rpl16 | M8 | 27 | −565.310154 | 0.000009183 | 5 K 0.966 *; 42 I 0.966 * |
| M7 | 25 | −576.908351 | |||
| rpl20 | M8 | 27 | −531.912203 | <0.000000001 | 76 H 0.998 **; 80 G 0.980 * |
| M7 | 25 | −574.476753 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, J.; Choi, M.-J.; Kim, S.-H.; Choi, H.-J.; Kim, S.-C. Plastome Characterization and Phylogenomic Analysis Yield New Insights into the Evolutionary Relationships among the Species of the Subgenus Bryocles (Hosta; Asparagaceae) in East Asia. Plants 2021, 10, 1980. https://doi.org/10.3390/plants10101980
Yang J, Choi M-J, Kim S-H, Choi H-J, Kim S-C. Plastome Characterization and Phylogenomic Analysis Yield New Insights into the Evolutionary Relationships among the Species of the Subgenus Bryocles (Hosta; Asparagaceae) in East Asia. Plants. 2021; 10(10):1980. https://doi.org/10.3390/plants10101980
Chicago/Turabian StyleYang, JiYoung, Mi-Jung Choi, Seon-Hee Kim, Hyeok-Jae Choi, and Seung-Chul Kim. 2021. "Plastome Characterization and Phylogenomic Analysis Yield New Insights into the Evolutionary Relationships among the Species of the Subgenus Bryocles (Hosta; Asparagaceae) in East Asia" Plants 10, no. 10: 1980. https://doi.org/10.3390/plants10101980