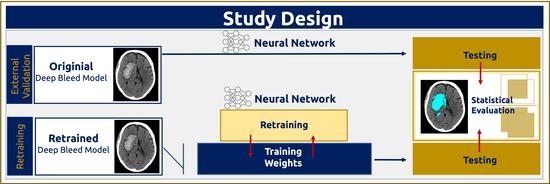
External Validation and Retraining of DeepBleed: The First Open-Source 3D Deep Learning Network for the Segmentation of Spontaneous Intracerebral and Intraventricular Hemorrhage
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Population
2.2. Image Acquisition and Manual Segmentation
2.3. Preprocessing and Postprocessing
2.4. Model Retraining
2.5. Model Testing
2.6. Code Availability
2.7. Statistical Analysis
3. Results
3.1. Demographics and Characteristics of the Study Cohort
3.2. Model Retraining and Testing
3.3. Analysis of Factors Influencing the Model Performance
3.4. Volume and Segmentation Agreement Analysis
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Feigin, V.L.; Stark, B.A.; Johnson, C.O.; Roth, G.A.; Bisignano, C.; Abady, G.G.; Abbasifard, M.; Abbasi-Kangevari, M.; Abd-Allah, F.; Abedi, V.; et al. Global, regional, and national burden of stroke and its risk factors, 1990–2019: A systematic analysis for the Global Burden of Disease Study 2019. Lancet Neurol. 2021, 20, 795–820. [Google Scholar] [CrossRef] [PubMed]
- Caplan, L.R. Intracerebral haemorrhage. Lancet 1992, 339, 656–658. [Google Scholar] [CrossRef] [PubMed]
- Qureshi, A.I.; Mendelow, A.D.; Hanley, D.F. Intracerebral haemorrhage. Lancet 2009, 373, 1632–1644. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pinho, J.; Costa, A.S.; Araújo, J.M.; Amorim, J.M.; Ferreira, C. Intracerebral hemorrhage outcome: A comprehensive update. J. Neurol. Sci. 2019, 398, 54–66. [Google Scholar] [CrossRef]
- Hemphill, J.C., III; Greenberg, S.M.; Anderson, C.S.; Becker, K.; Bendok, B.R.; Cushman, M.; Fung, G.L.; Goldstein, J.N.; Macdonald, R.L.; Mitchell, P.H.; et al. Guidelines for the Management of Spontaneous Intracerebral Hemorrhage: A Guideline for Healthcare Professionals from the American Heart Association/American Stroke Association. Stroke 2015, 46, 2032–2060. [Google Scholar] [CrossRef] [Green Version]
- Anderson, C.S.; Huang, Y.; Wang, J.G.; Arima, H.; Neal, B.; Peng, B.; Heeley, E.; Skulina, C.; Parsons, M.W.; Kim, J.S.; et al. Intensive blood pressure reduction in acute cerebral haemorrhage trial (INTERACT): A randomised pilot trial. Lancet Neurol. 2008, 7, 391–399. [Google Scholar] [CrossRef]
- Garg, R.K.; Liebling, S.M.; Maas, M.B.; Nemeth, A.J.; Russell, E.J.; Naidech, A.M. Blood pressure reduction, decreased diffusion on MRI, and outcomes after intracerebral hemorrhage. Stroke 2012, 43, 67–71. [Google Scholar] [CrossRef]
- Morgan, T.; Zuccarello, M.; Narayan, R.; Keyl, P.; Lane, K.; Hanley, D. Preliminary findings of the minimally-invasive surgery plus rtPA for intracerebral hemorrhage evacuation (MISTIE) clinical trial. Acta Neurochir. Suppl. 2008, 105, 147–151. [Google Scholar]
- Webb, A.J.; Ullman, N.L.; Morgan, T.C.; Muschelli, J.; Kornbluth, J.; Awad, I.A.; Mayo, S.; Rosenblum, M.; Ziai, W.; Zuccarrello, M.; et al. Accuracy of the ABC/2 Score for Intracerebral Hemorrhage: Systematic Review and Analysis of MISTIE, CLEAR-IVH, and CLEAR III. Stroke 2015, 46, 2470–2476. [Google Scholar] [CrossRef] [Green Version]
- Delcourt, C.; Carcel, C.; Zheng, D.; Sato, S.; Arima, H.; Bhaskar, S.; Janin, P.; Salman, R.A.-S.; Cao, Y.; Zhang, S.; et al. Comparison of ABC methods with computerized estimates of intracerebral hemorrhage volume: The INTERACT2 study. Cerebrovasc. Dis. Extra 2019, 9, 148–154. [Google Scholar] [CrossRef]
- Wang, T.; Song, N.; Liu, L.; Zhu, Z.; Chen, B.; Yang, W.; Chen, Z. Efficiency of a deep learning-based artificial intelligence diagnostic system in spontaneous intracerebral hemorrhage volume measurement. BMC Med. Imaging 2021, 21, 125. [Google Scholar] [CrossRef]
- Sharrock, M.F.; Mould, W.A.; Ali, H.; Hildreth, M.; Awad, I.A.; Hanley, D.F.; Muschelli, J. 3D Deep Neural Network Segmentation of Intracerebral Hemorrhage: Development and Validation for Clinical Trials. Neuroinformatics 2021, 19, 403–415. [Google Scholar] [CrossRef]
- Hanley, D.F.; Thompson, R.E.; Morgan, T.C.; Ullman, N.; Mould, W.A.; Carhuapoma, J.; Kase, C.; Ziai, W.; Thompson, C.B.; Yenokyan, G.; et al. Safety and efficacy of minimally invasive surgery plus alteplase in intracerebral haemorrhage evacuation (MISTIE): A randomised, controlled, open-label, phase 2 trial. Lancet Neurol. 2016, 15, 1228–1237. [Google Scholar] [CrossRef] [Green Version]
- Hanley, D.F.; Thompson, R.E.; Rosenblum, M.; Yenokyan, G.; Lane, K.; McBee, N.; Mayo, S.W.; Bistran-Hall, A.J.; Gandhi, D.; Mould, W.A.; et al. Efficacy and safety of minimally invasive surgery with thrombolysis in intracerebral haemorrhage evacuation (MISTIE III): A randomised, controlled, open-label, blinded endpoint phase 3 trial. Lancet 2019, 393, 1021–1032. [Google Scholar] [CrossRef] [Green Version]
- Yu, A.C.; Mohajer, B.; Eng, J. External validation of deep learning algorithms for radiologic diagnosis: A systematic review. Radiol. Artif. Intell. 2022, 4, e210064. [Google Scholar] [CrossRef]
- Hanley, D. Minimally Invasive Surgery Plus Rt-PA for ICH Evacuation Phase III (MISTIE III). Available online: https://clinicaltrials.gov/ct2/show/study/NCT01827046 (accessed on 28 January 2022).
- Falcone, G.J.; Biffi, A.; Brouwers, H.B.; Anderson, C.D.; Battey, T.W.K.; Ayres, A.; Vashkevich, A.; Schwab, K.; Rost, N.S.; Goldstein, J.N.; et al. Predictors of hematoma volume in deep and lobar supratentorial intracerebral hemorrhage. JAMA Neurol. 2013, 70, 988–994. [Google Scholar] [CrossRef] [Green Version]
- Chen, R.; Wang, X.; Anderson, C.S.; Ronbinson, T.; Lavados, P.M.; Lindley, R.I.; Chalmers, J.; Delcourt, C.; The INTERACT Investigators. Infratentorial intracerebral hemorrhage: Relation of location to outcome. Stroke 2019, 50, 1257–1259. [Google Scholar] [CrossRef]
- Yushkevich, P.A.; Piven, J.; Hazlett, H.C.; Smith, R.G.; Ho, S.; Gee, J.C.; Gerig, G. User-guided 3D active contour segmentation of anatomical structures: Significantly improved efficiency and reliability. Neuroimage 2006, 31, 1116–1128. [Google Scholar] [CrossRef] [Green Version]
- Harris, C.; Millman, K.; van der Walt, S.; Gommers, R.; Virtanen, P.; Cournapeau, D.; Smith Kern, R.; Picus, M.; Hoyer, S.; van Kerkwijk, M.H.; et al. Array programming with NumPy. Nature 2020, 585, 357–362. [Google Scholar] [CrossRef]
- Smith, S.M. Fast robust automated brain extraction. Hum. Brain Mapp. 2002, 17, 143–155. [Google Scholar] [CrossRef]
- Avants, B.B.; Tustison, N.; Song, G. Advanced normalization tools (ANTS). Insight J. 2009, 2, 1–35. [Google Scholar]
- Rorden, C.; Bonilha, L.; Fridriksson, J.; Bender, B.; Karnath, H.O. Age-specific CT and MRI templates for spatial normalization. Neuroimage 2012, 61, 957–965. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kingma, D.P.; Ba, J. Adam: A method for stochastic optimization. arXiv 2014, arXiv:14126980. [Google Scholar]
- Dice, L.R. Measures of the amount of ecologic association between species. Ecology 1945, 26, 297–302. [Google Scholar] [CrossRef]
- Van Rossum, G.; Drake, F.L. Python 3 Reference Manual; CreateSpace: Scotts Valley, CA, USA, 2009. [Google Scholar]
- McCarthy, P.; Cottaar, M.; Webster, M.; Fitzgibbon, S.; Craig, M. fslpy (3.10.0). 2022. Available online: https://git.fmrib.ox.ac.uk/fsl/fslpy/ (accessed on 9 May 2023).
- GraphPad. Available online: www.graphpad.com (accessed on 9 May 2023).
- Wickham, H.; Averick, M.; Bryan, J.; Chang, W.; McGowan, L.D.A.; François, R.; Grolemund, G.; Hayes, A.; Henry, L.; Hester, J.; et al. Welcome to the tidyverse. J. Open Source Softw. 2019, 4, 1686. [Google Scholar] [CrossRef] [Green Version]
- Muschelli, J.; Sweeney, E.M.; Ullman, N.L.; Vespa, P.; Hanley, D.F.; Crainiceanu, C.M. PItcHPERFeCT: Primary intracranial hemorrhage probability estimation using random forests on CT. NeuroImage Clin. 2017, 14, 379–390. [Google Scholar] [CrossRef]
- Morotti, A.; Goldstein, J.N. Anticoagulant-associated intracerebral hemorrhage. Brain Hemorrhages 2020, 1, 89–94. [Google Scholar] [CrossRef]
- Gerner, S.T.; Kuramatsu, J.B.; Sembill, J.A.; Sprügel, M.I.; Hagen, M.; Knappe, R.U.; Endres, M.; Haeusler, K.G.; Sobesky, J.; Schurig, J.; et al. Characteristics in Non–Vitamin K Antagonist Oral Anticoagulant–Related Intracerebral Hemorrhage. Stroke 2019, 50, 1392–1402. [Google Scholar] [CrossRef]
- Isensee, F.; Jaeger, P.F.; Kohl, S.A.; Petersen, J.; Maier-Hein, K.H. nnU-Net: A self-configuring method for deep learning-based biomedical image segmentation. Nat. Methods 2021, 18, 203–211. [Google Scholar] [CrossRef]
- Isensee, F.; Jäger, P.F.; Kohl, S.A.; Petersen, J.; Maier-Hein, K.H. Automated design of deep learning methods for biomedical image segmentation. arXiv 2019, arXiv:190408128. [Google Scholar]
- Zhao, X.; Chen, K.; Wu, G.; Zhang, G.; Zhou, X.; Lv, C.; Wu, S.; Chen, Y.; Xie, G.; Yao, Z. Deep learning shows good reliability for automatic segmentation and volume measurement of brain hemorrhage, intraventricular extension, and peripheral edema. Eur. Radiol. 2021, 31, 5012–5020. [Google Scholar] [CrossRef]
- Patel, A.; Leemput SCvd Prokop, M.; Ginneken, B.V.; Manniesing, R. Image Level Training and Prediction: Intracranial Hemorrhage Identification in 3D Non-Contrast CT. IEEE Access 2019, 7, 92355–92364. [Google Scholar] [CrossRef]
- Ironside, N.; Chen, C.-J.; Ding, D.; Mayer, S.A.; Connolly, E.S., Jr. Perihematomal edema after spontaneous intracerebral hemorrhage. Stroke 2019, 50, 1626–1633. [Google Scholar] [CrossRef]
- Dhar, R.; Falcone, G.J.; Chen, Y.; Hamzehloo, A.; Kirsch, E.P.; Noche, R.B.; Roth, K.; Acosta, J.; Ruiz, A.; Phuah, C.-L.; et al. Deep Learning for Automated Measurement of Hemorrhage and Perihematomal Edema in Supratentorial Intracerebral Hemorrhage. Stroke 2020, 51, 648–651. [Google Scholar] [CrossRef]
- Yu, N.; Yu, H.; Li, H.; Ma, N.; Hu, C.; Wang, J. A robust deep learning segmentation method for hematoma volumetric detection in intracerebral hemorrhage. Stroke 2022, 53, 167–176. [Google Scholar] [CrossRef]
- Lin, L.; Dou, Q.; Jin, Y.-M.; Zhou, G.-Q.; Tang, Y.-Q.; Chen, W.-L.; Su, B.-A.; Liu, F.; Tao, C.-J.; Jiang, N.; et al. Deep learning for automated contouring of primary tumor volumes by MRI for nasopharyngeal carcinoma. Radiology 2019, 291, 677–686. [Google Scholar] [CrossRef]
- Rudie, J.D.; Weiss, D.A.; Saluja, R.; Rauschecker, A.M.; Wang, J.; Sugrue, L.; Bakas, S.; Colby, J.B. Multi-disease segmentation of gliomas and white matter hyperintensities in the BraTS data using a 3D convolutional neural network. Front. Comput. Neurosci. 2019, 13, 84. [Google Scholar] [CrossRef] [Green Version]
- Scherer, M.; Cordes, J.; Younsi, A.; Sahin, Y.-A.; Götz, M.; Möhlenbrunch, M.; Stock, C.; Bösel, J.; Unterbreg, A.; Maier-Hein, K.; et al. Development and Validation of an Automatic Segmentation Algorithm for Quantification of Intracerebral Hemorrhage. Stroke 2016, 47, 2776–2782. [Google Scholar] [CrossRef] [Green Version]
- Koo, T.K.; Li, M.Y. A Guideline of Selecting and Reporting Intraclass Correlation Coefficients for Reliability Research. J. Chiropr. Med. 2016, 15, 155–163. [Google Scholar] [CrossRef] [Green Version]
- Cicchetti, D.V. Guidelines, Criteria, and Rules of Thumb for Evaluating Normed and Standardized Assessment Instruments in Psychology. Psychol. Assess. 1994, 6, 284–290. [Google Scholar] [CrossRef]
- Hanley, D. Clot Lysis: Evaluating Accelerated Resolution of Intraventricular Hemorrhage Phase III (CLEAR-III). 2013. Available online: https://clinicaltrials.gov/ct2/show/NCT00784134 (accessed on 28 January 2022).
- Hanley, D.F.; Lane, K.; McBee, N.; Ziai, W.; Tuhrim, S.; Lees, K.R.; Dawson, J.; Gandhi, D.; Ullman, N.; Mould, W.A.; et al. Thrombolytic removal of intraventricular haemorrhage in treatment of severe stroke: Results of the randomised, multicentre, multiregion, placebo-controlled CLEAR III trial. Lancet 2017, 389, 603–611. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- MIND. Artemis in the Removal of Intracerebral Hemoorrhage. Available online: https://clinicaltrials.gov/ct2/show/NCT03342664 (accessed on 28 January 2022).
- Taha, A.A.; Hanbury, A. Metrics for evaluating 3D medical image segmentation: Analysis, selection, and tool. BMC Med. Imaging 2015, 15, 29. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hemphill, J.C., III; Bonovich, D.C.; Besmertis, L.; Manley, G.T.; Johnston, S.C. The ICH score: A simple, reliable grading scale for intracerebral hemorrhage. Stroke 2001, 32, 891–897. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hill, M.D.; Silver, F.L.; Austin, P.C.; Tu, J.V. Rate of stroke recurrence in patients with primary intracerebral hemorrhage. Stroke 2000, 31, 123–127. [Google Scholar] [CrossRef] [Green Version]
- Hallevi, H.; Albright, K.C.; Aronowski, J.; Barreto, A.D.; Martin-Schild, S.; Khaja, A.M.; Gonzales, N.R.; Illoh, K.; Noser, E.A.; Grotta, J.C. Intraventricular hemorrhage: Anatomic relationships and clinical implications. Neurology 2008, 70, 848–852. [Google Scholar] [CrossRef] [Green Version]
- Cao, H.; Dell’Orco, A. DeepBleed Retrained Weights. Dataset Zenodo. Available online: https://doi.org/10.5281/zenodo.7616199 (accessed on 9 May 2023). [CrossRef]
- Cao, H.; Dell’Orco, A. DeepBleed Code. Code Gitub. Available online: https://github.com/orangepepermint/retraindeepbleed (accessed on 9 May 2023).






| Variable | Training Cohort (n = 100) | Validation Cohort (n = 20) | Test Cohort (n = 920) | p-Value | |
|---|---|---|---|---|---|
| Age (years), mean ± SD | 70.5 ± 13.1 | 68 ± 13.4 | 69.6 ± 14.2 | 0.84 1 | |
| Sex, n (%) | |||||
| Male | 41 (41) | 11 (55) | 516 (56) | ||
| Female | 59 (59) | 9 (45) | 404 (44) | ||
| NIHSS score, median (IQR) | 7.5 (10) | 10 (10) | 7 (5) | 0.70 1 | |
| GCS score, median (IQR) | 13 (7) | 14 (4) | 13 (8) | 0.11 2 | |
| Symptom onset to imaging (hours), median (IQR) | 4.3 (13.6) | 3.9 (7.6) | 4.23 (12.8) | 0.89 2 | |
| ICH location, n (%) | |||||
| Lobar | 44 (44) | 6 (30) | 338 (36.7) | ||
| Deep | 46 (46) | 7 (35) | 455 (49.5) | ||
| Brainstem | 3 (3) | 3 (15) | 41 (4.5) | ||
| Cerebellum | 7 (7) | 4 (20) | 86 (9.3) | ||
| ICH + IVH volume (ml), mean ± SD | 83.8 ± 47.2 | 56.4 ± 89.7 | 76.5 ± 44.9 | 0.67 2 | |
| ICH volume (ml), mean ± SD | 27.7 ± 30.2 | 34.2 ± 30.9 | 44.1 ± 14.2 | 0.30 2 | |
| IVH volume (ml), mean ± SD | 61.1 ± 49.1 | 22.2 ± 77.1 | 34.5 ± 42.5 | 0.64 2 | |
| Metric | All Locations | Deep | Lobar | Brainstem | Cerebellum |
|---|---|---|---|---|---|
| OM | |||||
| DSC | 0.84 (0.73, 0.88) | 0.86 (0.80, 0.89) | 0.84 (0.78, 0.89) | 0.71 (0.46, 0.78) | 0.48 (0.23, 0.64) |
| Sensitivity | 0.79 (0.65, 0.86) | 0.85 (0.79, 0.91) | 0.80, (0.70, 0.87) | 0.58 (0.38, 0.74) | 0.34 (0.13, 0.49) |
| PPV | 0.93 (0.85, 0.97) | 0.91 (0.85, 0.95) | 0.99 (0.85, 0.97) | 0.88 (0.76, 0.94) | 0.94 (0.76, 0.99) |
| RM | |||||
| DSC | 0.83 (0.74, 0.88) | 0.87 (0.81, 0.90) | 0.83 (0.72, 0.88) | 0.77 (0.57, 0.83) | 0.79 (0.65, 0.84) |
| Sensitivity | 0.80 (0.69, 0.87) | 0.85 (0.79, 0.91) | 0.79 (0.63, 0.88) | 0.72 (0.57, 0.79) | 0.75 (0.59, 0.84) |
| PPV | 0.91 (0.84, 0.95) | 0.91 (0.85, 0.95) | 0.92 (0.63, 0.88) | 0.87 (0.77, 0.94) | 0.88 (0.79, 0.94) |
| t1 OM vs. RM (padj-value) | |||||
| DSC | −5.9 (0.001) | 1.64 (ns) | 4.57 (0.001) | 1.90 (ns) | 12.94 (0.001) |
| Sensitivity | 1.45 (ns) | 3.05 (0.036) | 4.03 (0.001) | 3.33 (0.03) | 16.49 (0.001) |
| PPV | −7.23 (0.001) | 0.12 (ns) | 2.33 (ns) | 0.02 (ns) | 0.30 (ns) |
| OM | RM | |||||
|---|---|---|---|---|---|---|
| Parameter | Slope | SD | p-Value | Slope | SD | p-Value |
| 0.75 | 0.01 | <0.001 | 0.78 | 0.01 | <0.001 | |
| Location (in respect to deep location) | ||||||
| Lobar | −0.04 | 0.01 | <0.01 | −0.06 | 0.01 | <0.001 |
| Brainstem | −0.20 | 0.03 | <0.001 | −0.18 | 0.03 | <0.001 |
| Cerebellum | −0.32 | 0.02 | <0.001 | −0.08 | 0.02 | <0.001 |
| Volume (mm3) | 0.00 | 0.00 | <0.001 | 0.00 | 0.00 | <0.001 |
| IVH Presence | 0.02 | 0.01 | 0.17 | 0.02 | 0.01 | 0.15 |
| Center (in respect to Berlin, DE) | ||||||
| Hamburg, DE | 0.003 | 0.01 | 0.81 | −0.02 | 0.013 | 0.09 |
| Pavia, IT | 0.008 | 0.02 | 0.73 | −0.01 | 0.023 | 0.66 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cao, H.; Morotti, A.; Mazzacane, F.; Desser, D.; Schlunk, F.; Güttler, C.; Kniep, H.; Penzkofer, T.; Fiehler, J.; Hanning, U.; et al. External Validation and Retraining of DeepBleed: The First Open-Source 3D Deep Learning Network for the Segmentation of Spontaneous Intracerebral and Intraventricular Hemorrhage. J. Clin. Med. 2023, 12, 4005. https://doi.org/10.3390/jcm12124005
Cao H, Morotti A, Mazzacane F, Desser D, Schlunk F, Güttler C, Kniep H, Penzkofer T, Fiehler J, Hanning U, et al. External Validation and Retraining of DeepBleed: The First Open-Source 3D Deep Learning Network for the Segmentation of Spontaneous Intracerebral and Intraventricular Hemorrhage. Journal of Clinical Medicine. 2023; 12(12):4005. https://doi.org/10.3390/jcm12124005
Chicago/Turabian StyleCao, Haoyin, Andrea Morotti, Federico Mazzacane, Dmitriy Desser, Frieder Schlunk, Christopher Güttler, Helge Kniep, Tobias Penzkofer, Jens Fiehler, Uta Hanning, and et al. 2023. "External Validation and Retraining of DeepBleed: The First Open-Source 3D Deep Learning Network for the Segmentation of Spontaneous Intracerebral and Intraventricular Hemorrhage" Journal of Clinical Medicine 12, no. 12: 4005. https://doi.org/10.3390/jcm12124005