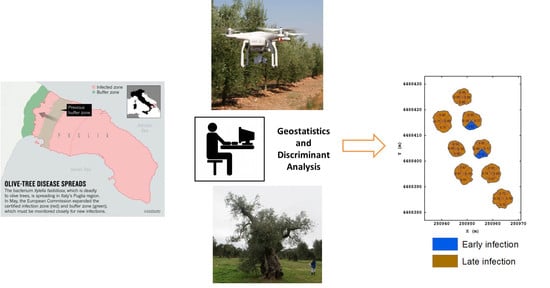
Semi-Automatic Method for Early Detection of Xylella fastidiosa in Olive Trees Using UAV Multispectral Imagery and Geostatistical-Discriminant Analysis
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Site Description
2.2. Variables
2.3. Methodology
- Extraction of canopies for each field from UAV images.
- ✓
- Pre-processing:
- To create a composite multispectral image from the individual spectral bands, the procedure of layer stacking has been applied, so individual image bands to have the same extent (no. of rows and columns).
- To enhance specific information about the landscape, which cannot easily be seen with a natural color image, as stress and vigor of vegetation, the false color composite has been applied. In this case, to emphasize the status of the plant, the following bands have been combined: Red → Red, Red-Edge → Green, Green → Blue, (Figure 4a).
- ✓
- Supervised Classification:
- Supervised classification was applied, which involves the use of training area data considered representative of the land cover types of the study areas. In this case three labels were identified: soil, canopy and shadow.As classifier algorithm, the maximum likelihood was used, which is based on maximizing the likelihood that the observed values follow a normal distribution (Figure 4b).Figure 4. (a) An example, Oria Field: 4 layer stack at false colors. (b) An example, Oria Field: thematic map of classification.Figure 4. (a) An example, Oria Field: 4 layer stack at false colors. (b) An example, Oria Field: thematic map of classification.
- To smooth the boundaries of small areas located near each other, or to aggregate these areas, a morphological filter [31,32,33], which uses the fundamental operations of erosion and dilation, was preferred to fill gaps in the contour lines. Subsequently, a format conversion has been applied to convert labelled raster image into vector data to extract closed spatial features (polygons) from the classification.
- ✓
- Exporting to GIS Environment:
- Each closed polygon, representing an individual plant, was then imported into GIS environment and an editing procedure was applied to generate a multipolygon product (Figure 5). This process allowed to modify the vertices of the selected spatial feature, to fill eventual holes in the polygon and/or to cut some parts.
- Each polygon was further subdivided into four quadrants (North, East, South, and West), to which to refer both visual and radiometric measurements (common support). The quadrants were generated by a procedure that has been implemented in C# language with ArcObjects libraries to be integrated into the ArcGIS environment. This procedure is based on segments that join the centroid of each polygon with the four points at 45°, 135°, 225°, 315° (defined above the horizon), to split the polygon into the north, east, south, and west sectors of the crown, respectively.
- Each shapefiles of quadrants has been imported in geostatistical environment.
- 2.
- Change of support: polygon cokriging.
2.4. Construction of the Prediction Model: Statistical Analyses
- Selection of predictors. The first step in the statistical analysis was preliminary to determine which variables (bands) were significantly related to the response class variable (disease severity level), coded as shown above. For this purpose, a stepwise discriminant analysis was performed, which is a regression technique aimed to select a subset of the quantitative variables (predictors) for use in discriminating between the classes. The variables are chosen to enter or to leave the regression model on the basis of the significance level of an F test from an analysis of covariance, where the variables already chosen act as covariates and the variable under consideration is the response variable [39]. The analysis was performed using the STEPDISC procedure of SAS software in stepwise mode [39], and the significance levels for a variable to enter the subset and to stay in the subset were set to 0.1.
- Check of multivariate normality. The next step for implementing discriminant analysis was to check the multivariate normality in each of the two severity classes of Xfp symptoms, since the estimation of misclassification probabilities requires the assumption of multivariate normality. Since the condition of normality for each of the 16 quantitative variables is a necessary, but not sufficient, condition for multi-normality, first, the assumption of univariate normality was checked with three tests (Kolmogorov–Smirnov, Cramer–von Mises, and Anderson–Darling) [40].
- 3.
- Testing the sensitivity of multi-band data to the infection severity level. Univariate (ANOVA) and multivariate (MANOVA) analyses of variance were carried out on the UAV data, the former to test the hypothesis that the class means for each quantitative variable (band reflectivity) were equal, whereas the latter to compare multivariate class means across several variables. Four multivariate statistical tests were used: Wilks’ lambda, Pillai’s trace, Hotelling–Lawley trace, and Roy’s maximum root [42,43,44,45,46,47].
- 4.
- Determination of the parametric discriminant model. Assuming that each severity class had a multivariate normal distribution, firstly, a parametric method was performed, aimed at developing a discriminant mathematical function or classification, which best separated between the two classes of severity [44]. The classification criterion can be a linear function, assuming the same variance–covariance matrix of responses across the severity classes, or quadratic, assuming each class with a unique variance structure. A chi-square test of equal variance was then performed [43]. Using Bayes theorem, the posterior probability of each observation belonging to each class was calculated, taking into account the prior probabilities of the classes [45,46]. Each observation was placed in the class to which it had the highest posterior probability to belong. Therefore, as the final product of the model, for each observation the most probable class was assigned together with its posterior probability, which can be considered as a measure of the uncertainty associated with such assignation or prediction.
- 5.
- Determination of non-parametric discriminant model. Since multivariate normality might not be fully satisfied, a non-parametric approach was also estimated to be compared with the parametric one. Non-parametric discriminant methods are based on non-parametric estimates of class-specific probability densities. The non-parametric kernel method uses a fixed radius (r) and a specified kernel (k), which can be uniform, normal, Epanechnikov, biweight or triweight, to calculate the kernel density in each class [44]. The value of r and type of kernel, called smoothing parameters, determine the degree of irregularity in the estimate of the density function. Small values of r produce jagged density estimates, whereas large values produce smoother density estimates. Therefore, for each type of kernel, several (ten on average) r values were tested, and the optimal set of the smoothing parameters, which minimizes the error rates, was chosen.
- 6.
- Comparison between the two models. The performance of each model was evaluated using cross-validation [45,46]. Cross-validation uses n-1 out of n observations to determine the discriminant function for the classification of the one observation left out. This is repeated for each of the n observations. The error-rate estimates were calculated by counting the number of misclassified observations; the class-specific error-count estimate was determined as the proportion of misclassified observations in the class. [47,48,49]. The overall error rate was calculated as a weighted average of the individual class-specific error-rate estimates by using the prior probabilities as the weights [50,51,52,53,54].
- 7.
- Graphical display of infection status. Canonical discriminant analysis was also performed to extract one (number of classes (2) minus 1) linear combination of the quantitative variables, called canonical variable, which best revealed the differences between the classes and had the highest possible multiple correlation with the classes. The standardized canonical coefficients were estimated, which express the partial contribution of each quantitative variable (band) to the canonical variable, and were then used to interpret its meaning.
- 8.
- Prediction phase. The better classification model was then used in the prediction phase, by determining the more likely severity class for an independent data set not used in the previous phase of construction of the model. In particular, eight plants in the Torchiarolo field, for which the visual surveys were missing, were used for severity class prediction on August 2019. The estimated posterior probability of the predicted class was also provided as a measure of prediction uncertainty.
3. Results
3.1. Geostatistical Analysis
3.2. Statistical Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Saponari, M.; Giampetruzzi, A.; Loconsole, G.; Boscia, D.; Saldarelli, P. Xylella fastidiosa in olive in Apulia: Where we stand. Phytopathology 2019, 109, 175–186. [Google Scholar] [CrossRef] [Green Version]
- Frem, M.; Chapman, D.; Fucilli, V.; Choueiri, E.; El Moujabber, M.; La Notte, P.; Nigro, F. Xylella fastidiosa invasion of new countries in Europe, the Middle East and North Africa: Ranking the potential exposure scenarios. NeoBiota 2020, 59, 77–97. [Google Scholar] [CrossRef]
- Calderón, R.; Navas-Cortés, J.A.; Lucena, C.; Zarco-Tejada, P.J. High-resolution airborne hyperspectral and thermal imagery for early detection of Verticillium wilt of olive using fluorescence, temperature and narrow-band spectral indices. Remote Sens. Environ. 2013, 139, 231–245. [Google Scholar] [CrossRef]
- Navas-Cortés, J.A.; Landa, B.B.; Mercado-Blanco, J.; Trapero-Casas, J.L.; Rodríguez-Jurado, D.; Jiménez-Díaz, R.M. Spatiotemporal analysis of spread of infections by Verticillium dahliae pathotypes within a high tree density olive orchard in Southern Spain. Phytopathology 2008, 98, 167–180. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Poblete, T.; Camino, C.; Beck, P.S.A.; Hornero, A.; Kattenborn, T.; Saponari, M.; Boscia, D.; Navas-Cortes, J.A.; Zarco-Tejada, P.J. Detection of Xylella fastidiosa infection symptoms with airborne multispectral and thermal imagery: Assessing bandset reduction performance from hyperspectral analysis. ISPRS J. Photogramm. Remote Sens. 2020, 162, 27–40. [Google Scholar] [CrossRef]
- Chester, K.S. Plant disease losses: Their appraisal and interpretation. In Plant Disease Reporter Supplement 193; Biodiversity Heritage Library: Washington, DC, USA, 1950; pp. 189–362. [Google Scholar]
- Cobb, N.A. Contribution to an economic knowledge of the Australian rusts (Uredinae). Agric. Gazt. N. S. W. 1892, 3, 44–48. [Google Scholar]
- Bock, C.H.; Poole, G.H.; Parker, P.E.; Gottwald, T.R. Plant disease severity estimated visually, by digital photography and image analysis, and by hyperspectral imaging. CRC. Crit. Rev. Plant Sci. 2010, 29, 59–107. [Google Scholar] [CrossRef]
- Kumar, A.; Lee, W.S.; Ehsani, R.J.; Albrigo, L.G.; Yang, C.; Mangan, R.L. Citrus greening disease detection using aerial hyperspectral and multispectral imaging techniques. J. Appl. Remote Sens. 2012, 6. [Google Scholar] [CrossRef]
- Li, X.; Lee, W.S.; Li, M.; Ehsani, R.; Mishra, A.R.; Yang, C.; Mangan, R.L. Spectral difference analysis and airborne imaging classification for citrus greening infected trees. Comput. Electron. Agric. 2012, 83, 32–46. [Google Scholar] [CrossRef]
- Usha, K.; Singh, B. Potential applications of remote sensing in horticulture-A review. Sci. Hortic. 2013, 153, 71–83. [Google Scholar] [CrossRef]
- Hornero, A.; Hernández-Clemente, R.; North, P.R.J.; Beck, P.S.A.; Boscia, D.; Navas-Cortes, J.A.; Zarco-Tejada, P.J. Monitoring the incidence of Xylella fastidiosa infection in olive orchards using ground-based evaluations, airborne imaging spectroscopy and Sentinel-2 time series through 3-D radiative transfer modelling. Remote Sens. Environ. 2020, 236, 111480. [Google Scholar] [CrossRef]
- Santesteban, L.G.; Di Gennaro, S.F.; Herrero-Langreo, A.; Miranda, C.; Royo, J.B.; Matese, A. High-resolution UAV-based thermal imaging to estimate the instantaneous and seasonal variability of plant water status within a vineyard. Agric. Water Manag. 2017, 183, 49–59. [Google Scholar] [CrossRef]
- Barbedo, J.G.A. A Review on the use of unmanned aerial vehicles and imaging sensors for monitoring and assessing plant stresses. Drones 2019, 3, 40. [Google Scholar] [CrossRef] [Green Version]
- Zhang, C.; Kovacs, J.M. The application of small unmanned aerial systems for precision agriculture: A review. Precis. Agric. 2012, 13, 693–712. [Google Scholar] [CrossRef]
- Zarco-Tejada, P.J.; Camino, C.; Beck, P.S.A.; Calderon, R.; Hornero, A.; Hernández-Clemente, R.; Kattenborn, T.; Montes-Borrego, M.; Susca, L.; Morelli, M.; et al. Previsual symptoms of Xylella fastidiosa infection revealed in spectral plant-trait alterations. Nat. Plants 2018, 4, 432–439. [Google Scholar] [CrossRef] [PubMed]
- Horler, D.N.H.; Dockray, M.; Barber, J. The red edge of plant leaf reflectance. Int. J. Remote Sens. 1983, 4, 273–288. [Google Scholar] [CrossRef]
- Otsu, K.; Pla, M.; Vayreda, J.; Brotons, L. Calibrating the severity of forest defoliation by pine processionary moth with Landsat and UAV imagery. Sensors 2018, 18, 3278. [Google Scholar] [CrossRef] [Green Version]
- Thenkabail, A.; Lyon, P.; Huete, J. Hyperspectral Remote Sensing of Vegetation, 1st ed.; Thenkabail, A., Lyon, P., Eds.; CRC Press: Boca Raton, FL, USA, 2011; Chapter 1; pp. 3–36. [Google Scholar] [CrossRef]
- Delwiche, S.R.; Kim, M.S. Hyperspectral imaging for detection of scab in wheat. In Environmental and Industrial Sensing: Biological Quality and Precision Agriculture II; SPIE: Boston, MA, USA, 2000. [Google Scholar] [CrossRef]
- Qin, J.; Burks, T.F.; Kim, M.S.; Chao, K.; Ritenour, M.A. Citrus canker detection using hyperspectral reflectance imaging and PCA-based image classification method. Sens. Instrum. Food Qual. Saf. 2008, 2, 168–177. [Google Scholar] [CrossRef]
- Delalieux, S.; van Aardt, J.; Keulemans, W.; Schrevens, E.; Coppin, P. Detection of biotic stress (Venturia inaequalis) in apple trees using hyperspectral data: Non-parametric statistical approaches and physiological implications. Eur. J. Agron. 2007, 27, 130–143. [Google Scholar] [CrossRef]
- Moshou, D.; Bravo, C.; West, J.; Wahlen, S.; McCartney, A.; Ramon, H. Automatic detection of “yellow rust” in wheat using reflectance measurements and neural networks. Comput. Electron. Agric. 2004, 44, 173–188. [Google Scholar] [CrossRef]
- Rumpf, T.; Mahlein, A.K.; Steiner, U.; Oerke, E.C.; Dehne, H.W.; Plümer, L. Early detection and classification of plant diseases with Support Vector Machines based on hyperspectral reflectance. Comput. Electron. Agric. 2010, 74, 91–99. [Google Scholar] [CrossRef]
- Camps-Valls, G.; Bruzzone, L. Kernel-based methods for hyperspectral image classification. IEEE Trans. Geosci. Remote Sens. 2005, 43, 1351–1362. [Google Scholar] [CrossRef]
- Olea, R.A. Geostatistical Glossary and Multilingual Dictionary; Oxford University Press: New York, NY, USA, 1991; p. 192. [Google Scholar]
- Castrignanò, A.; Buttafuoco, G. Data processing: Chapter 3. In Agricultural Internet of Things and Decision Support for Precision Smart Farming, 1st ed.; Castrignanò, A., Buttafuoco, G., Khosla, R., Mouazen, A.M., Moshou, D., Naud, O., Eds.; Academic Press: Cambridge, MA, USA, 2020; pp. 139–182. [Google Scholar]
- Gotway, C.A.; Young, L.J. Combining incompatible spatial data. J. Am. Stat. Assoc. 2002, 97, 632–648. [Google Scholar] [CrossRef] [Green Version]
- Matheron, G. Principles of geostatistics. Econ. Geol. 1963, 58, 1246–1266. [Google Scholar] [CrossRef]
- Castrignanò, A.; Quarto, R.; Venezia, A.; Buttafuoco, G. A comparison between mixed support kriging and block cokriging for modelling and combining spatial data with different support. Precis. Agric. 2019, 20, 193–213. [Google Scholar] [CrossRef]
- Lantuéjoul, C.; Serra, J. M-Filters. In Proceedings of the IEEE International Conference on Acoustics, Speech and Signal Processing ICASSP’82, Paris, France, 3–5 May 1982; pp. 2063–2066. [Google Scholar] [CrossRef]
- Serra, J. Image Analysis and Mathematical Morphology; Academic Press Inc.: Cambridge, MA, USA, 1983. [Google Scholar]
- Sternberg, S.R. Biomedical image processing. IEEE Comput. 1983, 16, 22–34. [Google Scholar] [CrossRef]
- Diacono, M.; Castrignanò, A.; Vitti, C.; Stellacci, A.M.; Marino, L.; Cocozza, C.; De Benedetto, D.; Troccoli, A.; Rubino, P.; Ventrella, D. An approach for assessing the effects of site-specific fertilization on crop growth and yield of durum wheat in organic agriculture. Precis. Agric. 2014, 15, 479–498. [Google Scholar] [CrossRef]
- Landrum, C.; Castrignanó, A.; Zourarakis, D.; Mueller, T. Assessing the time stability of soil moisture patterns using statistical and geostatistical approaches. Agric. Water Manag. 2016, 177, 118–127. [Google Scholar] [CrossRef]
- Goovaerts, P. Geostatistics for Natural Resources Evaluation; Oxford University Press: New York, NY, USA, 1997; p. 483. [Google Scholar]
- Wackernagel, H. Multivariate Geostatistics: An Introduction with Applications; Springer Nature: London, UK, 2003; ISBN 13 9783540441427. [Google Scholar]
- Castrignanò, A.; Belmonte, A.; Antelmi, I.; Quarto, R.; Quarto, F.; Shaddad, S.; Sion, V.; Muolo, M.R.; Ranieri, N.A.; Gadaleta, G.; et al. A geostatistical fusion approach using UAV data for probabilistic estimation of Xylella fastidiosa subsp. pauca infection in olive trees. STOTEN 2020, 752. [Google Scholar] [CrossRef]
- Journel, A.G.; Huijbregts, C.J. Mining Geostatistics; Academic Press: London, UK, 1978; 600p. [Google Scholar]
- Goovaerts, P. Factorial kriging analysis: A useful tool for exploring the structure of multivariate spatial soil information. J. Soil Sci. 1992, 43, 597–619. [Google Scholar] [CrossRef]
- Castrignanò, A.; Giugliarini, L.; Risaliti, R.; Martinelli, N. Study of spatial relationships among some soil physico-chemical properties of a field in central Italy using multivariate geostatistics. Geoderma 2000, 97, 39–60. [Google Scholar] [CrossRef]
- Castrignanò, A.; Buttafuoco, G.; Quarto, R.; Vitti, C.; Langella, G.; Terribile, F.; Venezia, A. A Combined Approach of Sensor Data Fusion and Multivariate Geostatistics for Delineation of Homogeneous Zones in an Agricultural Field. Sensors 2017, 17, 2794. [Google Scholar] [CrossRef] [PubMed]
- Chilès, J.P.; Delfiner, P. Geostatistics: Modeling Spatial Uncertainty, 2nd ed.; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2012. [Google Scholar] [CrossRef]
- Klecka, W.R. Discriminant Analysis; Sage University Paper Series; Sage Publications: Beverly Hills, CA, USA; London, UK, 1980; pp. 7–19. [Google Scholar]
- Evans, J.L.; Drew, J.H.; Leemis, L.M. The distribution of the Kolmogorov-Smirnov, Cramer-von Mises, and Anderson-Darling test Statistics for Exponential populations with Estimated Parameters. Commun. Stat. Simul. Comput. 2008, 37, 1396–1421. [Google Scholar] [CrossRef]
- Blom, G. Statistical Estimates and Transformed Beta Variables; John Wiley & Sons Inc.: New York, NY, USA, 1958. [Google Scholar]
- Morrison, D.F. Multivariate Statistical Methods, 3rd ed.; McGraw-Hill Inc.: New York, NY, USA, 1990; pp. 1–85. [Google Scholar]
- Rinaldi, M.; Castrignanò, A.; De Benedetto, D.; Sollitto, D.; Ruggieri, S.; Garofalo, P.; Santoro, F.; Figorito, B.; Gualano, S.; Tamborrino, R. Discrimination of tomato plants under different irrigation regimes: Analysis of hyperspectral sensor data. Environmetrics 2015. [CrossRef]
- Epanechnikov, V.A. Non-parametric estimation of a multivariate probability density. Theor. Probab. Appl. 1969, 14, 153–158. [Google Scholar] [CrossRef]
- Lachenbruch, P.A.; Mickey, M.R. Estimation of error rates in discriminant analysis. Technometrics 1968, 10, 1–11. [Google Scholar] [CrossRef]
- Fukunaga, K.; Kessel, D.L. Nonparametric Bayes error estimation using unclassified samples. IEEE Trans. Inform. Theor. 1973, 19, 434–440. [Google Scholar] [CrossRef]
- Glick, N. Additive Estimators for probabilities of correct classification. Pattern Recogn. 1978, 10, 211–222. [Google Scholar] [CrossRef]
- Hora, S.C.; Wilcox, J.B. Estimation of error rates in several-population discriminant analysis. J. Mark. Res. 1982, 19, 57–61. [Google Scholar] [CrossRef]
- Sims, D.A.; Gamon, J.A. Relationships between leaf pigment content and spectral reflectance across a wide range of species, leaf structures and developmental stages. Remote Sens. Environ. 2002, 81, 337–354. [Google Scholar] [CrossRef]
- Girke, T. Programming in R-Manuals. Available online: http://manuals.bioinformatics.ucr.edu/home/programming-in-r (accessed on 20 December 2020).
- Diggle, P.J.; Ribeiro, R.P. Model-Based Geostatistics; Springer Science Business Media LLC: New York, NY, USA, 2007; ISBN 9780387485362. [Google Scholar]
- QGIS. A Free and Open Source Geographic Information System. Available online: https://qgis.org/it/site/ (accessed on 20 December 2020).











| Field | Type of Model | Range (m) | Spatial Variance Explained (%) |
|---|---|---|---|
| Re | Nugget effect | - | 32 |
| Cardinal sinus | 18.64 | 68 | |
| Fella | Nugget effect | - | 37 |
| Cubic | 2.42 | 21 | |
| Cardinal sinus | 31.18 | 42 | |
| Torchiarolo | Nugget effect | - | 54 |
| Cardinal sinus | 22.23 | 37 | |
| Spherical | 88.62 | 9 |
| Variable | D* | W-Qu* | A-Qu* | |||
|---|---|---|---|---|---|---|
| Statistics | Probability | Statistic | Probability | Statistic | Probability | |
| GREEN | 0.281019 | <0.0100 | 105.9703 | <0.0050 | 585.016 | <0.0050 |
| RED | 0.21957 | <0.0100 | 56.25436 | <0.0050 | 331.2315 | <0.0050 |
| RED-EDGE | 0.164006 | <0.0100 | 17.94999 | <0.0050 | 123.1129 | <0.0050 |
| NIR | 0.066529 | <0.0100 | 2.045712 | <0.0050 | 10.74146 | <0.0050 |
| GREEN_std | 0.442217 | <0.0100 | 157.4172 | <0.0050 | 830.784 | <0.0050 |
| RED_std | 0.391301 | <0.0100 | 126.0456 | <0.0050 | 728.0533 | <0.0050 |
| RED-EDGE_std | 0.370405 | <0.0100 | 111.0805 | <0.0050 | 552.6192 | <0.0050 |
| NIR_std | 0.281603 | <0.0100 | 65.14581 | <0.0050 | 323.1493 | <0.0050 |
| Variable | F Value | Probability |
|---|---|---|
| R_GREEN | 134.09 | <0.0001 |
| R_RED | 24.07 | <0.0001 |
| R_RED-EDGE | 174.09 | <0.0001 |
| R_NIR | 106.90 | <0.0001 |
| R_GREEN_std | 50.57 | <0.0001 |
| R_RED_std | 0.84 | 0.3580 |
| R_RED-EDGE_std | 90.59 | <0.0001 |
| R_NIR_std | 16.86 | <0.0001 |
| Test | Statistic Value | F Value | Probability |
|---|---|---|---|
| Wilks’s lambda | 0.931 | 37.74 | <0.0001 |
| Pillai’s trace | 0.069 | 37.74 | <0.0001 |
| Hotelling–Lawley’s trace | 0.074 | 37.74 | <0.0001 |
| Roy’s maximum root | 0.074 | 37.74 | <0.0001 |
| Ground Truth | |||||
| 0 | 1 | Total # of classified samples | User’s accuracy | ||
| Classification Results | 0 | 1405 | 498 | 1903 | 0.74 |
| 1 | 968 | 1224 | 2192 | 0.56 | |
| Total #of ground truth samples | 2373 | 1722 | |||
| Producer’s accuracy | 0.59 | 0.71 | 0.64 | ||
| Class | 0 | 1 | Average |
|---|---|---|---|
| Error Rate | 0.41 | 0.29 | 0.36 |
| Ground Truth | |||||
| 0 | 1 | Total # of classified samples | User’s accuracy | ||
| Classification Results | 0 | 1835 | 730 | 2565 | 0.72 |
| 1 | 538 | 992 | 1530 | 0.65 | |
| Total #of ground truth samples | 2373 | 1722 | |||
| Producer’s accuracy | 0.77 | 0.58 | 0.69 | ||
| Class | 0 | 1 | Average |
|---|---|---|---|
| Error rate | 0.23 | 0.42 | 0.31 |
| Variable | Coefficients |
|---|---|
| R_GREEN | 0.034 |
| R_RED | 0.652 |
| R_RED-EDGE | 0.885 |
| R_NIR | −0.247 |
| R_GREEN_std | 0.171 |
| R_RED_std | −0.689 |
| R_RED-EDGE_std | 0.364 |
| R_NIR_std | −0.154 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Castrignanò, A.; Belmonte, A.; Antelmi, I.; Quarto, R.; Quarto, F.; Shaddad, S.; Sion, V.; Muolo, M.R.; Ranieri, N.A.; Gadaleta, G.; et al. Semi-Automatic Method for Early Detection of Xylella fastidiosa in Olive Trees Using UAV Multispectral Imagery and Geostatistical-Discriminant Analysis. Remote Sens. 2021, 13, 14. https://doi.org/10.3390/rs13010014
Castrignanò A, Belmonte A, Antelmi I, Quarto R, Quarto F, Shaddad S, Sion V, Muolo MR, Ranieri NA, Gadaleta G, et al. Semi-Automatic Method for Early Detection of Xylella fastidiosa in Olive Trees Using UAV Multispectral Imagery and Geostatistical-Discriminant Analysis. Remote Sensing. 2021; 13(1):14. https://doi.org/10.3390/rs13010014
Chicago/Turabian StyleCastrignanò, Annamaria, Antonella Belmonte, Ilaria Antelmi, Ruggiero Quarto, Francesco Quarto, Sameh Shaddad, Valentina Sion, Maria Rita Muolo, Nicola A. Ranieri, Giovanni Gadaleta, and et al. 2021. "Semi-Automatic Method for Early Detection of Xylella fastidiosa in Olive Trees Using UAV Multispectral Imagery and Geostatistical-Discriminant Analysis" Remote Sensing 13, no. 1: 14. https://doi.org/10.3390/rs13010014