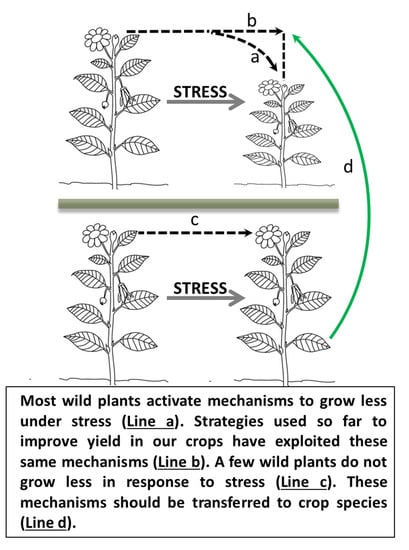
It’s Hard to Avoid Avoidance: Uncoupling the Evolutionary Connection between Plant Growth, Productivity and Stress “Tolerance”
Abstract
:1. Preamble
2. Avoidance—The Basis of Plant Abiotic Stress Tolerance
3. Survival and Avoidance of Injury and Death Are Negatively Correlated with Growth, Biomass and Productivity
4. Plant Growth versus Survival and Productivity
5. The Energy Limitation Myth
6. A New Conceptual Framework
7. Genomics, Bioinformatics and Integration
8. Thinking around the Dogmas—Searching for Answers where the Problems Are
Funding
Acknowledgments
Conflicts of Interest
References
- Giovannini, I.; Altiero, T.; Guidetti, R.; Rebecchi, L. Will the Antarctic tardigrade Acutuncus antarcticus be able to withstand environmental stresses related to global climate change? J. Exp. Biol. 2018, 221. [Google Scholar] [CrossRef] [PubMed]
- Maas, E.V.; Hoffman, G.J. Crop salt tolerance–current assessment. J. Irrig. Drain. Div. 1977, 102, 115–134. [Google Scholar]
- Umezawa, T.; Fujita, M.; Fujita, Y.; Yamaguchi-Shinozaki, K.; Shinozaki, K. Engineering drought tolerance in plants: Discovering and tailoring genes to unlock the future. Curr. Opin. Biotechnol. 2006, 17, 113–122. [Google Scholar] [CrossRef] [PubMed]
- Maggio, A.; Zhu, J.K.; Hasegawa, P.M.; Bressan, R.A. Osmogenetics: Aristotle to Arabidopsis. Plant Cell 2006, 18, 1542–1557. [Google Scholar] [CrossRef] [PubMed]
- Pardo, J.M. Biotechnology of water and salinity stress tolerance. Curr. Opin. Biotechnol. 2010, 21, 185–196. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yoshida, T.; Fujita, Y.; Maruyama, K.; Mogami, J.; Todaka, D.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Four Arabidopsis AREB/ABF transcription factors function predominantly in gene expression downstream of SnRK2 kinases in abscisic acid signalling in response to osmotic stress: Transcriptional regulation in SnRK2 signalling. Plant Cell Environ. 2015, 38, 35–49. [Google Scholar] [CrossRef] [PubMed]
- Gao, S.; Gao, J.; Zhu, X.; Song, Y.; Li, Z.; Ren, G.; Zhou, X.; Kuai, B. ABF2, ABF3, and ABF4 Promote ABA-Mediated Chlorophyll Degradation and Leaf Senescence by Transcriptional Activation of Chlorophyll Catabolic Genes and Senescence-Associated Genes in Arabidopsis. Mol. Plant 2016, 9, 1272–1285. [Google Scholar] [CrossRef] [PubMed]
- Thalmann, M.; Pazmino, D.; Seung, D.; Horrer, D.; Nigro, A.; Meier, T.; Kölling, K.; Pfeifhofer, H.W.; Zeeman, S.C.; Santelia, D. Regulation of Leaf Starch Degradation by Abscisic Acid Is Important for Osmotic Stress Tolerance in Plants. Plant Cell 2016, 28, 1860–1878. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ma, Q.-J.; Sun, M.-H.; Lu, J.; Liu, Y.-J.; Hu, D.-G.; Hao, Y.-J. Transcription Factor AREB2 Is Involved in Soluble Sugar Accumulation by Activating Sugar Transporter and Amylase Genes. Plant Physiol. 2017, 174, 2348–2362. [Google Scholar] [CrossRef] [PubMed]
- Maggio, A.; Miyazaki, S.; Veronese, P.; Fujita, T.; Ibeas, J.I.; Damsz, B.; Narasimhan, M.L.; Hasegawa, P.M.; Joly, R.J.; Bressan, R.A. Does proline accumulation play an active role in stress-induced growth reduction? Plant J. 2002, 31, 699–712. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, C.; Liu, Y.; Li, S.-S.; Han, G.-Z. Insights into the Origin and Evolution of the Plant Hormone Signaling Machinery. Plant Physiol. 2015, 167, 872–886. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, Y.; Chan, Z.; Gao, J.; Xing, L.; Cao, M.; Yu, C.; Hu, Y.; You, J.; Shi, H.; Zhu, Y.; et al. ABA receptor PYL9 promotes drought resistance and leaf senescence. Proc. Natl. Acad. Sci. USA 2012, 113, 1949–1954. [Google Scholar] [CrossRef] [PubMed]
- Tischer, S.V.; Wunschel, C.; Papacek, M.; Kleigrewe, K.; Hofmann, T.; Christmann, A.; Grill, E. Combinatorial interaction network of abscisic acid receptors and coreceptors from Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2017, 114, 10280–10285. [Google Scholar] [CrossRef] [PubMed]
- Blum, A. Drought resistance, water-use efficiency, and yield potential—Are they compatible, dissonant, or mutually exclusive? Aust. J. Agric. Res. 2005, 56, 1159. [Google Scholar] [CrossRef]
- Blum, A. Plant Breeding for Water-Limited Environments; Springer: New York, NY, USA, 2011; ISBN 978-1-4419-7490-7. [Google Scholar]
- Li, S.-L.; Xia, Y.-Z.; Sun, Z.-Q. Effects of cold-shock on the growth and flower bud differentiation of tomato seedlings under high temperature stress. Chin. J. Appl. Ecol. 2016, 27, 477–483. [Google Scholar] [CrossRef]
- Shavrukov, Y.; Kurishbayev, A.; Jatayev, S.; Shvidchenko, V.; Zotova, L.; Koekemoer, F.; de Groot, S.; Soole, K.; Langridge, P. Early Flowering as a Drought Escape Mechanism in Plants: How Can It Aid Wheat Production? Front. Plant Sci. 2017, 8. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, Y.; Chan, Z.; Xing, L.; Liu, X.; Hou, Y.-J.; Chinnusamy, V.; Wang, P.; Duan, C.; Zhu, J.-K. The unique mode of action of a divergent member of the ABA-receptor protein family in ABA and stress signaling. Cell Res. 2013, 23, 1380–1395. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Miao, C.; Xiao, L.; Hua, K.; Zou, C.; Zhao, Y.; Bressan, R.A.; Zhu, J.-K. Mutations in a subfamily of abscisic acid receptor genes promote rice growth and productivity. Proc. Natl. Acad. Sci. USA 2018, 115, 6058–6063. [Google Scholar] [CrossRef] [PubMed]
- Park, S.-Y.; Peterson, F.C.; Mosquna, A.; Yao, J.; Volkman, B.F.; Cutler, S.R. Agrochemical control of plant water use using engineered abscisic acid receptors. Nature 2015, 520, 545–548. [Google Scholar] [CrossRef] [PubMed]
- Cutler, S.R.; Rodriguez, P.L.; Finkelstein, R.R.; Abrams, S.R. Abscisic Acid: Emergence of a Core Signaling Network. Annu. Rev. Plant Biol. 2010, 61, 651–679. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Raghavendra, A.S.; Gonugunta, V.K.; Christmann, A.; Grill, E. ABA perception and signalling. Trends Plant Sci. 2010, 15, 395–401. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.-K. Abiotic Stress Signaling and Responses in Plants. Cell 2016, 167, 313–324. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Flowers, T.J. Improving crop salt tolerance. J. Exp. Bot. 2004, 55, 307–319. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fujii, H.; Chinnusamy, V.; Rodrigues, A.; Rubio, S.; Antoni, R.; Park, S.-Y.; Cutler, S.R.; Sheen, J.; Rodriguez, P.L.; Zhu, J.-K. In vitro reconstitution of an abscisic acid signalling pathway. Nature 2009, 462, 660–664. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ma, Y.; Szostkiewicz, I.; Korte, A.; Moes, D.; Yang, Y.; Christmann, A.; Grill, E. Regulators of PP2C Phosphatase Activity Function as Abscisic Acid Sensors. Science 2009. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.-K. Plant salt tolerance. Trends Plant Sci. 2001, 2, 66–71. [Google Scholar] [CrossRef]
- Inan, G.; Zhang, Q.; Li, P.; Wang, Z.; Cao, Z.; Zhang, H.; Zhang, C.; Quist, T.M.; Goodwin, S.M.; Zhu, J.; et al. Salt Cress. A Halophyte and Cryophyte Arabidopsis Relative Model System and Its Applicability to Molecular Genetic Analyses of Growth and Development of Extremophiles. Plant Phys. 2004, 135, 1718–1737. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gong, Q.; Li, P.; Ma, S.; Indu Rupassara, S.; Bohnert, H.J. Salinity stress adaptation competence in the extremophile Thellungiella halophila in comparison with its relative Arabidopsis thaliana: Salinity stress adaptation in T. halophila. Plant J. 2005, 44, 826–839. [Google Scholar] [CrossRef] [PubMed]
- Du, J.; Huang, Y.-P.; Xi, J.; Cao, M.-J.; Ni, W.-S.; Chen, X.; Zhu, J.-K.; Oliver, D.J.; Xiang, C.-B. Functional gene-mining for salt-tolerance genes with the power of Arabidopsis. Plant J. 2008, 56, 653–664. [Google Scholar] [CrossRef] [PubMed]
- Amtmann, A. Learning from Evolution: Thellungiella Generates New Knowledge on Essential and Critical Components of Abiotic Stress Tolerance in Plants. Mol. Plant 2009, 2, 3–12. [Google Scholar] [CrossRef] [PubMed]
- Oh, D.-H.; Dassanayake, M.; Haas, J.S.; Kropornika, A.; Wright, C.; d’Urzo, M.P.; Hong, H.; Ali, S.; Hernandez, A.; Lambert, G.M.; et al. Genome Structures and Halophyte-Specific Gene Expression of the Extremophile Thellungiella parvula in Comparison with Thellungiella salsuginea (Thellungiella halophila) and Arabidopsis. Plant Physiol. 2010, 154, 1040–1052. [Google Scholar] [CrossRef] [PubMed]
- Eshel, G.; Shaked, R.; Kazachkova, Y.; Khan, A.; Eppel, A.; Cisneros, A.; Acuna, T.; Gutterman, Y.; Tel-Zur, N.; Rachmilevitch, S.; et al. Anastatica hierochuntica, an Arabidopsis Desert Relative, Is Tolerant to Multiple Abiotic Stresses and Exhibits Species-Specific and Common Stress Tolerance Strategies with Its Halophytic Relative, Eutrema (Thellungiella) salsugineum. Front. Plant Sci. 2017, 7. [Google Scholar] [CrossRef] [PubMed]
- Ben-Romdhane, W.; Ben-Saad, R.; Meynard, D.; Zouari, N.; Mahjoub, A.; Fki, L.; Guiderdoni, E.; Al-Doss, A.; Hassairi, A. Overexpression of AlTMP2 gene from the halophyte grass Aeluropus littoralis in transgenic tobacco enhances tolerance to different abiotic stresses by improving membrane stability and deregulating some stress-related genes. Protoplasma 2018, 255, 1161–1177. [Google Scholar] [CrossRef] [PubMed]
- Orsini, F.; D’Urzo, M.P.; Inan, G.; Serra, S.; Oh, D.-H.; Mickelbart, M.V.; Consiglio, F.; Li, X.; Jeong, J.C.; Yun, D.-J.; et al. A comparative study of salt tolerance parameters in 11 wild relatives of Arabidopsis thaliana. J. Exp. Bot. 2010, 61, 3787–3798. [Google Scholar] [CrossRef] [PubMed]
- Dassanayake, M.; Oh, D.-H.; Haas, J.S.; Hernandez, A.; Hong, H.; Ali, S.; Yun, D.-J.; Bressan, R.A.; Zhu, J.-K.; Bohnert, H.J.; et al. The genome of the extremophile crucifer Thellungiella parvula. Nat. Genet. 2011, 43, 913–918. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.-J.; Zhang, Z.; Wang, J.-Y.; Oh, D.-H.; Dassanayake, M.; Liu, B.; Huang, Q.; Sun, H.-X.; Xia, R.; Wu, Y.; et al. Insights into salt tolerance from the genome of Thellungiella salsuginea. Proc. Natl. Acad. Sci. USA 2012, 109, 12219–12224. [Google Scholar] [CrossRef] [PubMed]
- Vera-Estrella, R.; Barkla, B.J.; García-Ramírez, L.; Pantoja, O. Salt Stress in Thellungiella halophila Activates Na+ Transport Mechanisms Required for Salinity Tolerance. Plant Phys. 2005, 139, 1507–1517. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.-L.; Chen, A.-P.; Zhong, N.-Q.; Liu, N.; Wu, X.-M.; Wang, F.; Yang, C.-L.; Romero, M.F.; Xia, G.-X. The Thellungiella salsuginea Tonoplast Aquaporin TsTIP1;2 Functions in Protection Against Multiple Abiotic Stresses. Plant Cell Physiol. 2014, 55, 148–161. [Google Scholar] [CrossRef] [PubMed]
- Marinho, G.S.; Holdt, S.L.; Birkeland, M.J.; Angelidaki, I. Commercial cultivation and bioremediation potential of sugar kelp, Saccharina latissima, in Danish waters. J. Appl. Phycol. 2015, 27, 1963–1973. [Google Scholar] [CrossRef]
- Ali, A.; Raddatz, N.; Aman, R.; Kim, S.; Park, H.C.; Jan, M.; Baek, D.; Khan, I.U.; Oh, D.-H.; Lee, S.Y.; et al. A Single Amino-Acid Substitution in the Sodium Transporter HKT1 Associated with Plant Salt Tolerance. Plant Physiol. 2016, 171, 2112–2126. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- An, D.; Chen, J.-G.; Gao, Y.-Q.; Li, X.; Chao, Z.-F.; Chen, Z.-R.; Li, Q.-Q.; Han, M.-L.; Wang, Y.-L.; Wang, Y.-F.; et al. AtHKT1 drives adaptation of Arabidopsis thaliana to salinity by reducing floral sodium content. PLOS Genet. 2017, 13, e1007086. [Google Scholar] [CrossRef] [PubMed]
- Queitsch, C.; Sangster, T.A.; Lindquist, S. Hsp90 as a capacitor of phenotypic variation. Nature 2002, 417, 618–624. [Google Scholar] [CrossRef] [PubMed]
- Ohno, S. Evolution by Gene Duplication; Springer-Verlag: Berlin, Germany; University of Michigan: Ann Arbor, MI, USA, 1970. [Google Scholar]
- Piatigorsky, J. The recruitment of crystallins: New functions precede gene duplication. Science 1991, 252, 1078–1079. [Google Scholar] [CrossRef] [PubMed]
- Des Marais, D.L.; Rausher, M.D. Escape from adaptive conflict after duplication in an anthocyanin pathway gene. Nature 2008, 454, 762–765. [Google Scholar] [CrossRef] [PubMed]
- Zhan, X.; Qian, B.; Cao, F.; Wu, W.; Yang, L.; Guan, Q.; Gu, X.; Wang, P.; Okusolubo, T.A.; Dunn, S.L.; et al. An Arabidopsis PWI and RRM motif-containing protein is critical for pre-mRNA splicing and ABA responses. Nat. Commun. 2015, 6. [Google Scholar] [CrossRef] [PubMed]
- Gu, J.; Xia, Z.; Luo, Y.; Jiang, X.; Qian, B.; Xie, H.; Zhu, J.-K.; Xiong, L.; Zhu, J.; Wang, Z.-Y. Spliceosomal protein U1A is involved in alternative splicing and salt stress tolerance in Arabidopsis thaliana. Nucleic Acids Res. 2018, 46, 1777–1792. [Google Scholar] [CrossRef] [PubMed]
- Levitt, J. Responses of Plants to Environmental Stresses; Academic Press: New York, NY, USA, 1972; p. 697. ISBN 0124455018. [Google Scholar]
- Bell, M.A.; Fischer, R.A.; Byerlee, D.; Sayre, K. Genetic and agronomic contributions to yield gains: A case study for wheat. Field Crops Res. 1995, 44, 55–65. [Google Scholar] [CrossRef]
- Evans, L.T.; Fischer, R.A. Yield Potential: Its Definition, Measurement, and Significance. Crop Sci. 1999, 39, 1544–1551. [Google Scholar] [CrossRef]
- Vinocur, B.; Altman, A. Recent advances in engineering plant tolerance to abiotic stress: Achievements and limitations. Curr. Opin. Biotechnol. 2005, 16, 123–132. [Google Scholar] [CrossRef] [PubMed]
- Boyer, J.S. Plant Productivity and Environment. Science 1982, 218, 443–448. [Google Scholar] [CrossRef] [PubMed]
- Zinselmeier, C.; Sun, Y.; Helentjaris, T.; Beatty, M.; Yang, S.; Smith, H.; Habben, J. The use of gene expression profiling to dissect the stress sensitivity of reproductive development in maize. Field Crops Res. 2002, 75, 111–121. [Google Scholar] [CrossRef]
- Miyazaki, S.; Fredricksen, M.; Hollis, K.C.; Poroyko, V.; Shepley, D.; Galbraith, D.W.; Long, S.P.; Bohnert, H.J. Transcript expression profiles of Arabidopsis thaliana grown under controlled conditions and open-air elevated concentrations of CO2 and of O3. Field Crops Res. 2004, 90, 47–59. [Google Scholar] [CrossRef]
- Ruggiero, B.; Koiwa, H.; Manabe, Y.; Quist, T.M.; Inan, G.; Saccardo, F.; Joly, R.J.; Hasegawa, P.M.; Bressan, R.A.; Maggio, A. Uncoupling the Effects of Abscisic Acid on Plant Growth and Water Relations. Analysis of sto1/nced3, an Abscisic Acid-Deficient but Salt Stress-Tolerant Mutant in Arabidopsis. Plant Physiol. 2004, 136, 3134–3147. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, Y.; Gao, J.; Im Kim, J.; Chen, K.; Bressan, R.A.; Zhu, J.-K. Control of Plant Water Use by ABA Induction of Senescence and Dormancy: An Overlooked Lesson from Evolution. Plant Cell Physiol. 2017, 58, 1319–1327. [Google Scholar] [CrossRef] [PubMed]
- Xiong, Y.; McCormack, M.; Li, L.; Hall, Q.; Xiang, C.; Sheen, J. Glucose–TOR signalling reprograms the transcriptome and activates meristems. Nature 2013, 496, 181–186. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, P.; Zhao, Y.; Li, Z.; Hsu, C.-C.; Liu, X.; Fu, L.; Hou, Y.-J.; Du, Y.; Xie, S.; Zhang, C.; et al. Reciprocal Regulation of the TOR Kinase and ABA Receptor Balances Plant Growth and Stress Response. Mol. Cell 2018, 69, 100–112.e6. [Google Scholar] [CrossRef] [PubMed]
- Tardieu, F.; Katerji, N.; Bethenod, O.; Zhang, J.; Davies, W.J. Maize stomatal conductance in the field: Its relationship with soil and plant water potentials, mechanical constraints and ABA concentration in the xylem sap. Plant Cell Environ. 1991, 14, 121–126. [Google Scholar] [CrossRef]
- Chaves, M.M. How Plants Cope with Water Stress in the Field? Photosynthesis and Growth. Ann. Bot. 2002, 89, 907–916. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- De Pascale, S.; Orsini, F.; Caputo, R.; Palermo, M.A.; Barbieri, G.; Maggio, A. Seasonal and multiannual effects of salinisation on tomato yield and fruit quality. Funct. Plant Biol. 2012, 39, 689–698. [Google Scholar] [CrossRef]
- Pardo, J.M.; Reddy, M.P.; Yang, S.; Maggio, A.; Huh, G.-H.; Matsumoto, T.; Coca, M.A.; Paino-D’Urzo, M.; Koiwa, H.; Yun, D.-J.; et al. Stress signaling through Ca2+/calmodulin-dependent protein phosphatase calcineurin mediates salt adaptation in plants. Proc. Natl. Acad. Sci. USA 1998, 95, 9681–9686. [Google Scholar] [CrossRef] [PubMed]
- Yobi, A.; Schlauch, K.A.; Perryman, B.; Oliver, M.J.; Cushman, J.C. Biomass Production, Nutritional and Mineral Content of Desiccation-Sensitive and Desiccation-Tolerant Species of Sporobolus under Multiple Irrigation Regimes. J. Agron. Crop Sci. 2013, 199, 309–320. [Google Scholar] [CrossRef]
- Xiao, L.; Yang, G.; Zhang, L.; Yang, X.; Zhao, S.; Ji, Z.; Zhou, Q.; Hu, M.; Wang, Y.; Chen, M.; et al. The resurrection genome of Boea hygrometrica: A blueprint for survival of dehydration. Proc. Natl. Acad. Sci. USA 2015, 112, 5833–5837. [Google Scholar] [CrossRef] [PubMed]
- Pnueli, L.; Hallak-Herr, E.; Rozenberg, M.; Cohen, M.; Goloubinoff, P.; Kaplan, A.; Mittler, R. Molecular and biochemical mechanisms associated with dormancy and drought tolerance in the desert legume Retama raetam. Plant J. 2002, 31, 319–330. [Google Scholar] [CrossRef] [PubMed]
- Dobzhansky, T. Biology, Molecular and Organismic. Am. Zool. 1964, 4, 443–452. [Google Scholar] [CrossRef] [PubMed]
- Bressan, R.A.; Zhu, J.-K.; Van Oosten, M.J.; Maggio, A.; Bohnert, H.J.; Chinnusamy, V. Epigenetics Connects the Genome to Its Environment. In Plant Breeding Reviews; Wiley-Blackwell: Hoboken, NJ, USA, 2014; Volume 38, pp. 69–142. ISBN 978-1-118-91686-5. [Google Scholar]
- Chinnusamy, V.; Gong, Z.; Zhu, J.-K. Nuclear RNA export and its importance in abiotic stress responses of plants. Curr. Top. Microbiol. Immunol. 2008, 326, 235–255. [Google Scholar] [PubMed]
- Ben Amor, B.; Wirth, S.; Merchan, F.; Laporte, P.; d’Aubenton-Carafa, Y.; Hirsch, J.; Maizel, A.; Mallory, A.; Lucas, A.; Deragon, J.M.; et al. Novel long non-protein coding RNAs involved in Arabidopsis differentiation and stress responses. Genome Res. 2009, 19, 57–69. [Google Scholar] [CrossRef] [PubMed]
- Doebley, J. The Genetics of Maize Evolution. Annu. Rev. Genet. 2004, 38, 37–59. [Google Scholar] [CrossRef] [PubMed]
- Chapman, M.A.; Pashley, C.H.; Wenzler, J.; Hvala, J.; Tang, S.; Knapp, S.J.; Burke, J.M. A Genomic Scan for Selection Reveals Candidates for Genes Involved in the Evolution of Cultivated Sunflower (Helianthus annuus). Plant Cell 2008, 20, 2931–2945. [Google Scholar] [CrossRef] [PubMed]
- Sang, T. Genes and Mutations Underlying Domestication Transitions in Grasses. Plant Physiol. 2009, 149, 63–70. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Monna, L.; Kitazawa, N.; Yoshino, R.; Suzuki, J.; Masuda, H.; Maehara, Y.; Tanji, M.; Sato, M.; Nasu, S.; Minobe, Y. Positional Cloning of Rice Semidwarfing Gene, sd-1: Rice “Green Revolution Gene” Encodes a Mutant Enzyme Involved in Gibberellin Synthesis. DNA Res. 2002, 9, 11–17. [Google Scholar] [CrossRef] [PubMed]
- Sasaki, A.; Ashikari, M.; Ueguchi-Tanaka, M.; Itoh, H.; Nishimura, A.; Swapan, D.; Ishiyama, K.; Saito, T.; Kobayashi, M.; Khush, G.S.; et al. A mutant gibberellin-synthesis gene in rice: Green revolution. Nature 2002, 416, 701–702. [Google Scholar] [CrossRef] [PubMed]
- Spielmeyer, W.; Ellis, M.H.; Chandler, P.M. Semidwarf (sd-1), “green revolution” rice, contains a defective gibberellin 20-oxidase gene. Proc. Natl. Acad. Sci. USA 2002, 99, 9043–9048. [Google Scholar] [CrossRef] [PubMed]
- Asano, K.; Yamasaki, M.; Takuno, S.; Miura, K.; Katagiri, S.; Ito, T.; Doi, K.; Wu, J.; Ebana, K.; Matsumoto, T.; et al. Artificial selection for a green revolution gene during japonica rice domestication. Proc. Natl. Acad. Sci. USA 2011, 108, 11034–11039. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ueguchi-Tanaka, M.; Ashikari, M.; Nakajima, M.; Itoh, H.; Katoh, E.; Kobayashi, M.; Chow, T.; Hsing, Y.C.; Kitano, H.; Yamaguchi, I.; et al. GIBBERELLIN INSENSITIVE DWARF1 encodes a soluble receptor for gibberellin. Nature 2005, 437, 693–698. [Google Scholar] [CrossRef] [PubMed]
- Matsuda, K.; Riazi, A. Stress-Induced Osmotic Adjustment in Growing Regions of Barley Leaves. Plant Physiol. 1981, 68, 571–576. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Glenn, E.P.; Brown, J.J.; Blumwald, E. Salt Tolerance and Crop Potential of Halophytes. Crit. Rev. Plant Sci. 1999, 18, 227–255. [Google Scholar] [CrossRef]
- Bressan, R.A.; Zhang, C.; Zhang, H.; Hasegawa, P.M.; Bohnert, H.J.; Zhu, J.-K. Learning from the Arabidopsis Experience. The Next Gene Search Paradigm. Plant Physiol. 2001, 127, 1354–1360. [Google Scholar] [CrossRef] [Green Version]
- Yang, Z.; Liu, J.; Tischer, S.V.; Christmann, A.; Windisch, W.; Schnyder, H.; Grill, E. Leveraging abscisic acid receptors for efficient water use in Arabidopsis. Proc. Natl. Acad. Sci. USA 2016, 113, 6791–6796. [Google Scholar] [CrossRef] [PubMed]
- Bressan, R.A.; Nelson, D.E.; Iraki, N.M.; La Rosa, C.P.; Singh, N.K.; Hasegawa, P.M.; Carpita, N.C. Reduced cell expansion and changes in cell walls of plant cells adapted to NaCl. In Environmental Injury to Plants; Katterman, F., Ed.; Academic Press: San Diego, CA, USA, 1990; pp. 137–171. [Google Scholar]
- Zhu, X.-G.; Long, S.P.; Ort, D.R. What is the maximum efficiency with which photosynthesis can convert solar energy into biomass? Curr. Opin. Biotechnol. 2008, 19, 153–159. [Google Scholar] [CrossRef] [PubMed]
- Slattery, R.A.; Ort, D.R. Photosynthetic Energy Conversion Efficiency: Setting a Baseline for Gauging Future Improvements in Important Food and Biofuel Crops. Plant Physiol. 2015, 168, 383–392. [Google Scholar] [CrossRef] [PubMed]
- Briggs, L.J.; Shantz, H.L. The Wilting Coefficient and Its Indirect Determination. Bot. Gaz. 1912, 53, 20–37. [Google Scholar] [CrossRef]
- Shantz, H.L. Soil moisture in relation to the growth of plants. J. Am. Soc. Agron. 1925, 17, 705–711. [Google Scholar] [CrossRef]
- Wiens, D.; Slaton, M.R. The mechanism of background extinction. Biol. J. Linn. Soc. 2012, 105, 255–268. [Google Scholar] [CrossRef]
- Tollenaar, M.; Lee, E.A. Yield potential, yield stability and stress tolerance in maize. Field Crop. Res. 2002, 75, 161–169. [Google Scholar] [CrossRef]
- Zoebl, D. Is water productivity a useful concept in agricultural water management? Agric. Water Manag. 2006, 84, 265–273. [Google Scholar] [CrossRef]
- Rozema, J.; Flowers, T. Crops for a Salinized World. Science 2008, 322, 1478–1480. [Google Scholar] [CrossRef] [PubMed]
- Niazi, B.H.; Rozema, J.; Broekman, R.A.; Salim, M. Dynamics of Growth and Water Relations of Fodderbeet and Seabeet in Response to Salinity. J. Agron. Crop Sci. 2000, 184, 101–110. [Google Scholar] [CrossRef]
- Goel, V.L.; Behl, H.M. Genetic selection and improvement of hard wood tree species for fuelwood production on sodic soil with particular reference to Prosopis juliflora. Biomass Bioenergy 2001, 20, 9–15. [Google Scholar] [CrossRef]
- Abideen, Z.; Ansari, R.; Khan, M.A. Halophytes: Potential source of ligno-cellulosic biomass for ethanol production. Biomass Bioenergy 2011, 35, 1818–1822. [Google Scholar] [CrossRef]
- Adnan, M.Y.; Hussain, T.; Asrar, H.; Hameed, A.; Gul, B.; Nielsen, B.L.; Khan, M.A. Desmostachya bipinnata manages photosynthesis and oxidative stress at moderate salinity. Flora 2016, 225, 1–9. [Google Scholar] [CrossRef]
- Amtmann, A.; Bohnert, H.J.; Bressan, R.A. Abiotic Stress and Plant Genome Evolution. Search for New Models. Plant Physiol. 2005, 138, 127–130. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Feng, Z.; Zhang, B.; Ding, W.; Liu, X.; Yang, D.-L.; Wei, P.; Cao, F.; Zhu, S.; Zhang, F.; Mao, Y.; et al. Efficient genome editing in plants using a CRISPR/Cas system. Cell Res. 2013, 23, 1229–1232. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kreps, J.A.; Wu, Y.; Chang, H.-S.; Zhu, T.; Wang, X.; Harper, J.F. Transcriptome Changes for Arabidopsis in Response to Salt, Osmotic, and Cold Stress. Plant Phys. 2002, 130, 2129–2141. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Oono, Y.; Seki, M.; Nanjo, T.; Narusaka, M.; Fujita, M.; Satoh, R.; Satou, M.; Sakurai, T.; Ishida, J.; Akiyama, K.; et al. Monitoring expression profiles of Arabidopsis gene expression during rehydration process after dehydration using ca. 7000 full-length cDNA microarray. Plant J. 2003, 34, 868–887. [Google Scholar] [CrossRef] [PubMed]
- Ma, S.; Bohnert, H.J. Integration of Arabidopsis thaliana stress-related transcript profiles, promoter structures, and cell-specific expression. Genome Biol. 2007, 8, R49. [Google Scholar] [CrossRef] [PubMed]
- Ma, S.; Gong, Q.; Bohnert, H.J. An Arabidopsis gene network based on the graphical Gaussian model. Genome Res. 2007, 17, 1614–1625. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fujii, H.; Verslues, P.E.; Zhu, J.-K. Arabidopsis decuple mutant reveals the importance of SnRK2 kinases in osmotic stress responses in vivo. Proc. Natl. Acad. Sci. USA 2011, 108, 1717–1722. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fujii, H.; Zhu, J.-K. Arabidopsis mutant deficient in 3 abscisic acid-activated protein kinases reveals critical roles in growth, reproduction, and stress. Proc. Natl. Acad. Sci. USA 2009, 106, 8380–8385. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fujita, Y.; Nakashima, K.; Yoshida, T.; Katagiri, T.; Kidokoro, S.; Kanamori, N.; Umezawa, T.; Fujita, M.; Maruyama, K.; Ishiyama, K.; et al. Three SnRK2 Protein Kinases are the Main Positive Regulators of Abscisic Acid Signaling in Response to Water Stress in Arabidopsis. Plant Cell Physiol. 2009, 50, 2123–2132. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gonzalez-Guzman, M.; Pizzio, G.A.; Antoni, R.; Vera-Sirera, F.; Merilo, E.; Bassel, G.W.; Fernández, M.A.; Holdsworth, M.J.; Perez-Amador, M.A.; Kollist, H.; et al. Arabidopsis PYR/PYL/RCAR Receptors Play a Major Role in Quantitative Regulation of Stomatal Aperture and Transcriptional Response to Abscisic Acid. Plant Cell 2012, 24, 2483–2496. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jia, Y.; Ding, Y.; Shi, Y.; Zhang, X.; Gong, Z.; Yang, S. The cbfs triple mutants reveal the essential functions of CBFs in cold acclimation and allow the definition of CBF regulons in Arabidopsis. New Phytol. 2016, 212, 345–353. [Google Scholar] [CrossRef] [PubMed]
- Nakashima, K.; Fujita, Y.; Kanamori, N.; Katagiri, T.; Umezawa, T.; Kidokoro, S.; Maruyama, K.; Yoshida, T.; Ishiyama, K.; Kobayashi, M.; et al. Three Arabidopsis SnRK2 Protein Kinases, SRK2D/SnRK2.2, SRK2E/SnRK2.6/OST1 and SRK2I/SnRK2.3, Involved in ABA Signaling are Essential for the Control of Seed Development and Dormancy. Plant Cell Physiol. 2009, 50, 1345–1363. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sakuraba, Y.; Kim, Y.-S.; Han, S.-H.; Lee, B.-D.; Paek, N.-C. The Arabidopsis Transcription Factor NAC016 Promotes Drought Stress Responses by Repressing AREB1 Transcription through a Trifurcate Feed-Forward Regulatory Loop Involving NAP. Plant Cell 2015, 27, 1771–1787. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, T.; Fujita, Y.; Sayama, H.; Kidokoro, S.; Maruyama, K.; Mizoi, J.; Shinozaki, K.; Yamaguchi-Shinozaki, K. AREB1, AREB2, and ABF3 are master transcription factors that cooperatively regulate ABRE-dependent ABA signaling involved in drought stress tolerance and require ABA for full activation. Plant J. 2010, 61, 672–685. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, C.; Zhang, Z.; Xie, S.; Si, T.; Li, Y.; Zhu, J.-K. Mutational Evidence for the Critical Role of CBF Genes in Cold Acclimation in Arabidopsis. Plant Physiol 2016, 00533. [Google Scholar] [CrossRef] [PubMed]
- Munns, R.; James, R.A.; Xu, B.; Athman, A.; Conn, S.J.; Jordans, C.; Byrt, C.S.; Hare, R.A.; Tyerman, S.D.; Tester, M.; et al. Wheat grain yield on saline soils is improved by an ancestral Na+ transporter gene. Nat. Biotechnol. 2012, 30, 360–364. [Google Scholar] [CrossRef] [PubMed]
- Arai-Sanoh, Y.; Takai, T.; Yoshinaga, S.; Nakano, H.; Kojima, M.; Sakakibara, H.; Kondo, M.; Uga, Y. Deep rooting conferred by DEEPER ROOTING 1 enhances rice yield in paddy fields. Sci. Rep. 2015, 4. [Google Scholar] [CrossRef] [PubMed]
- Bhaskara, G.B.; Nguyen, T.T.; Verslues, P.E. Unique Drought Resistance Functions of the Highly ABA-Induced Clade A Protein Phosphatase 2Cs. Plant Physiol. 2012, 160, 379–395. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cao, M.; Liu, X.; Zhang, Y.; Xue, X.; Zhou, X.E.; Melcher, K.; Gao, P.; Wang, F.; Zeng, L.; Zhao, Y.; et al. An ABA-mimicking ligand that reduces water loss and promotes drought resistance in plants. Cell Res. 2013, 23, 1043–1054. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cao, M.-J.; Zhang, Y.-L.; Liu, X.; Huang, H.; Zhou, X.E.; Wang, W.-L.; Zeng, A.; Zhao, C.-Z.; Si, T.; Du, J.; et al. Combining chemical and genetic approaches to increase drought resistance in plants. Nat. Commun. 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Dupeux, F.; Antoni, R.; Betz, K.; Santiago, J.; Gonzalez-Guzman, M.; Rodriguez, L.; Rubio, S.; Park, S.-Y.; Cutler, S.R.; Rodriguez, P.L.; et al. Modulation of Abscisic Acid Signaling in Vivo by an Engineered Receptor-Insensitive Protein Phosphatase Type 2C Allele. Plant Physiol. 2011, 156, 106–116. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kim, H.; Lee, K.; Hwang, H.; Bhatnagar, N.; Kim, D.-Y.; Yoon, I.S.; Byun, M.-O.; Kim, S.T.; Jung, K.-H.; Kim, B.-G. Overexpression of PYL5 in rice enhances drought tolerance, inhibits growth, and modulates gene expression. J. Exp. Bot. 2014, 65, 453–464. [Google Scholar] [CrossRef] [PubMed]
- Mosquna, A.; Peterson, F.C.; Park, S.-Y.; Lozano-Juste, J.; Volkman, B.F.; Cutler, S.R. Potent and selective activation of abscisic acid receptors in vivo by mutational stabilization of their agonist-bound conformation. Proc. Natl. Acad. Sci. USA 2011, 108, 20838–20843. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Okamoto, M.; Peterson, F.C.; Defries, A.; Park, S.-Y.; Endo, A.; Nambara, E.; Volkman, B.F.; Cutler, S.R. Activation of dimeric ABA receptors elicits guard cell closure, ABA-regulated gene expression, and drought tolerance. Proc. Natl. Acad. Sci. USA 2013, 110, 12132–12137. [Google Scholar] [CrossRef] [PubMed]
- Fujita, Y.; Fujita, M.; Satoh, R.; Maruyama, K.; Parvez, M.M.; Seki, M.; Hiratsu, K.; Ohme-Takagi, M.; Shinozaki, K.; et al. AREB1 Is a Transcription Activator of Novel ABRE-Dependent ABA Signaling That Enhances Drought Stress Tolerance in Arabidopsis. Plant Cell 2005, 17, 3470–3488. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Furihata, T.; Maruyama, K.; Fujita, Y.; Umezawa, T.; Yoshida, R.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Abscisic acid-dependent multisite phosphorylation regulates the activity of a transcription activator AREB1. Proc. Natl. Acad. Sci. USA 2006, 103, 1988–1993. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lawlor, D.W. Genetic engineering to improve plant performance under drought: Physiological evaluation of achievements, limitations, and possibilities. J. Exp. Bot. 2013, 64, 83–108. [Google Scholar] [CrossRef] [PubMed]
- Kudo, M.; Kidokoro, S.; Yoshida, T.; Mizoi, J.; Todaka, D.; Fernie, A.R.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Double overexpression of DREB and PIF transcription factors improves drought stress tolerance and cell elongation in transgenic plants. Plant Biotechnol. J. 2017, 15, 458–471. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.Z.; Creelman, R.A.; Zhu, J.K. From Laboratory to Field. Using Information from Arabidopsis to Engineer Salt, Cold, and Drought Tolerance in Crops. Plant Physiol. 2004, 135, 615–621. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Strabala, T.J.; Bednarek, S.Y.; Bertoni, G.; Amasino, R.M. Isolation and characterization of an ipt gene from the Ti plasmid Bo542. Mol. Gen. Genet. 1989, 216, 388–394. [Google Scholar] [CrossRef] [PubMed]
- Rivero, R.M.; Kojima, M.; Gepstein, A.; Sakakibara, H.; Mittler, R.; Gepstein, S.; Blumwald, E. Delayed leaf senescence induces extreme drought tolerance in a flowering plant. Proc. Natl. Acad. Sci. USA 2007, 104, 19631–19636. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nelson, D.E.; Repetti, P.P.; Adams, T.R.; Creelman, R.A.; Wu, J.; Warner, D.C.; Anstrom, D.C.; Bensen, R.J.; Castiglioni, P.P.; Donnarummo, M.G.; et al. Plant nuclear factor Y (NF-Y) B subunits confer drought tolerance and lead to improved corn yields on water-limited acres. Proc. Natl. Acad. Sci. USA 2007, 104, 16450–16455. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, Y.; Zhang, Z.; Gao, J.; Wang, P.; Hu, T.; Wang, Z.; Hou, Y.-J.; Wan, Y.; Liu, W.; Xie, S.; et al. Arabidopsis Duodecuple Mutant of PYL ABA Receptors Reveals PYL Repression of ABA-Independent SnRK2 Activity. Cell Rep. 2018, 23, 3340–3351.e5. [Google Scholar] [CrossRef] [PubMed]
- Castiglioni, P.; Warner, D.; Bensen, R.J.; Anstrom, D.C.; Harrison, J.; Stoecker, M.; Abad, M.; Kumar, G.; Salvador, S.; D’Ordine, R.; et al. Bacterial RNA Chaperones Confer Abiotic Stress Tolerance in Plants and Improved Grain Yield in Maize under Water-Limited Conditions. Plant Physiol. 2008, 147, 446–455. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Marris, E. Water: More crop per drop. Nature 2008, 45, 273–277. [Google Scholar] [CrossRef] [PubMed]
- Taji, T.; Ohsumi, C.; Iuchi, S.; Seki, M.; Kasuga, M.; Kobayashi, M.; Yamaguchi-Shinozaki, K.; Shinozaki, K. Important roles of drought- and cold-inducible genes for galactinol synthase in stress tolerance in Arabidopsis thaliana. Plant J. 2002, 29, 417–426. [Google Scholar] [CrossRef] [PubMed]
- Seki, M.; Umezawa, T.; Urano, K.; Shinozaki, K. Regulatory metabolic networks in drought stress responses. Curr. Opin. Plant Biol. 2007, 10, 296–302. [Google Scholar] [CrossRef] [PubMed]
- Park, S.; Lee, C.-M.; Doherty, C.J.; Gilmour, S.J.; Kim, Y.; Thomashow, M.F. Regulation of the Arabidopsis CBF regulon by a complex low-temperature regulatory network. Plant J. 2015, 82, 193–207. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Weng, J.-K.; Ye, M.; Li, B.; Noel, J.P. Co-evolution of Hormone Metabolism and Signaling Networks Expands Plant Adaptive Plasticity. Cell 2016, 166, 881–893. [Google Scholar] [CrossRef] [PubMed]
- Yuan, W.; Luo, X.; Li, Z.; Yang, W.; Wang, Y.; Liu, R.; Du, J.; He, Y. A cis cold memory element and a trans epigenome reader mediate Polycomb silencing of FLC by vernalization in Arabidopsis. Nat. Genet. 2016, 48, 1527–1534. [Google Scholar] [CrossRef] [PubMed]
- Geng, Y.; Wu, R.; Wee, C.W.; Xie, F.; Wei, X.; Chan, P.M.Y.; Tham, C.; Duan, L.; Dinneny, J.R. A Spatio-Temporal Understanding of Growth Regulation during the Salt Stress Response in Arabidopsis. Plant Cell 2013, 25, 2132–2154. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Takahashi, F.; Suzuki, T.; Osakabe, Y.; Betsuyaku, S.; Kondo, Y.; Dohmae, N.; Fukuda, H.; Yamaguchi-Shinozaki, K.; Shinozaki, K. A small peptide modulates stomatal control via abscisic acid in long-distance signalling. Nature 2018, 556, 235–238. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Xing, L.; Wang, X.; Hou, Y.-J.; Gao, J.; Wang, P.; Duan, C.-G.; Zhu, X.; Zhu, J.-K. The ABA Receptor PYL8 Promotes Lateral Root Growth by Enhancing MYB77-Dependent Transcription of Auxin-Responsive Genes. Sci. Signal. 2014, 7, ra53. [Google Scholar] [CrossRef] [PubMed]
- Scott, S.; Kranz, J.E.; Cole, J.; Lincecum, J.M.; Thompson, K.; Kelly, N.; Bostrom, A.; Theodoss, J.; Al-Nakhala, B.M.; Vieira, F.G.; et al. Design, power, and interpretation of studies in the standard murine model of ALS. Amyotroph. Lateral Scler. 2008, 9, 4–15. [Google Scholar] [CrossRef] [PubMed]
- Narasimhan, M.L.; Coca, M.A.; Jin, J.; Yamauchi, T.; Ito, Y.; Kadowaki, T.; Kim, K.K.; Pardo, J.M.; Damsz, B.; Hasegawa, P.M.; et al. Osmotin Is a Homolog of Mammalian Adiponectin and Controls Apoptosis in Yeast through a Homolog of Mammalian Adiponectin Receptor. Mol. Cell 2005, 17, 171–180. [Google Scholar] [CrossRef] [PubMed]
- Henderson, I.R.; Jacobsen, S.E. Epigenetic inheritance in plants. Nature 2007, 447, 418–424. [Google Scholar] [CrossRef] [PubMed]
- Sunkar, R.; Chinnusamy, V.; Zhu, J.; Zhu, J.-K. Small RNAs as big players in plant abiotic stress responses and nutrient deprivation. Trends Plant Sci. 2007, 12, 301–309. [Google Scholar] [CrossRef] [PubMed]
- Boyko, A.; Kovalchuk, I. Epigenetic control of plant stress response. Environ. Mol. Mutagen. 2008, 49, 61–72. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Purugganan, M.D.; Fuller, D.Q. The nature of selection during plant domestication. Nature 2009, 457, 843–848. [Google Scholar] [CrossRef] [PubMed]
- Keurentjes, J.J.; Koornneef, M.; Vreugdenhil, D. Quantitative genetics in the age of omics. Curr. Opin. Plant Biol. 2008, 11, 123–128. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pea, G.; Paulstephenraj, P.; Canè, M.A.; Sardaro, M.L.S.; Landi, P.; Morgante, M.; Porceddu, E.; Pè, M.E.; Frascaroli, E. Recombinant near-isogenic lines: A resource for the mendelization of heterotic QTL in maize. Mol. Genet. Genom. 2009, 281, 447–457. [Google Scholar] [CrossRef] [PubMed]
- Wilkinson, M.D.; Dumontier, M.; Aalbersberg, I.J.; Appleton, G.; Axton, M.; Baak, A.; Blomberg, N.; Boiten, J.-W.; da Silva Santos, L.B.; Bourne, P.E.; et al. The FAIR Guiding Principles for scientific data management and stewardship. Sci. Data 2016, 3, 160018. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rodríguez-Iglesias, A.; Rodríguez-González, A.; Irvine, A.G.; Sesma, A.; Urban, M.; Hammond-Kosack, K.E.; Wilkinson, M.D. Publishing FAIR Data: An Exemplar Methodology Utilizing PHI-Base. Front. Plant Sci. 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Van Bel, M.; Diels, T.; Vancaester, E.; Kreft, L.; Botzki, A.; Van de Peer, Y.; Coppens, F.; Vandepoele, K. PLAZA 4.0: An integrative resource for functional, evolutionary and comparative plant genomics. Nucleic Acids Res. 2018, 46, D1190–D1196. [Google Scholar] [CrossRef] [PubMed]
- Ziegler, G.R.; Hartsock, R.H.; Baxter, I. Zbrowse: An interactive GWAS results browser. PeerJ Comput. Sci. 2015, 1, e3. [Google Scholar] [CrossRef]
- Weigel, D.; Mott, R. The 1001 Genomes Project for Arabidopsis thaliana. Genome Biol. 2009, 10, 107. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.-K. Genetic Analysis of Plant Salt Tolerance Using Arabidopsis. Plant Physiol. 2000, 124, 941–948. [Google Scholar] [CrossRef] [PubMed]
- Miller, R.N.G.; Costa Alves, G.S.; Van Sluys, M.-A. Plant immunity: Unravelling the complexity of plant responses to biotic stresses. Ann. Bot. 2017, 119, 681–687. [Google Scholar] [CrossRef] [PubMed]
- Vaughan, D.A.; Institute, I.R.R. The Wild Relatives of Rice: A Genetic Resources Handbook; IRRI, International Rice Research Institute: Los Baños, Philippines, 1994; ISBN 978-971-22-0057-1. [Google Scholar]
- Ma, X.; Ding, Y.; Zhou, B.; Guo, W.; Lv, Y.; Zhu, X.; Zhang, T. QTL mapping in A-genome diploid Asiatic cotton and their congruence analysis with AD-genome tetraploid cotton in genus Gossypium. J. Genet. Genom. 2008, 35, 751–762. [Google Scholar] [CrossRef]
- Campos, H.; Cooper, M.; Habben, J.E.; Edmeades, G.O.; Schussler, J.R. Improving drought tolerance in maize: A view from industry. Field Crops Res. 2004, 90, 19–34. [Google Scholar] [CrossRef]
- Kromdijk, J.; Głowacka, K.; Leonelli, L.; Gabilly, S.T.; Iwai, M.; Niyogi, K.K.; Long, S.P. Improving photosynthesis and crop productivity by accelerating recovery from photoprotection. Science 2016, 354, 857–861. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Evans, L.T. Feeding the Ten Billion: Plants and Population Growth; Cambridge University Press: Cambridge, UK, 1998; ISBN 978-0-521-64685-7. [Google Scholar]
- Maggio, A.; Van Criekinge, T.; Malingreau, J.-P. Global food security: Assessing trends in view of guiding future EU policies. Foresight 2016, 18, 551–560. [Google Scholar] [CrossRef]
- McKersie, B. Planning for food security in a changing climate. J. Exp. Bot. 2015, 66, 3435–3450. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hardin, L.S. Bellagio 1969: The green revolution: Meetings that changed the world. Nature 2008, 455, 470–471. [Google Scholar] [CrossRef] [PubMed]
- Maggio, A.; Joly, R.J.; Hasegawa, P.M.; Bressan, R.A. Can the Quest for Drought Tolerant Crops Avoid Arabidopsis Any Longer? J. Crop Prod. 2003, 7, 99–129. [Google Scholar] [CrossRef]
- Flowers, T.J.; Colmer, T.D. Salinity tolerance in halophytes. New Phytol. 2008, 179, 945–963. [Google Scholar] [CrossRef] [PubMed]
- Taji, T.; Seki, M.; Satou, M.; Sakurai, T.; Kobayashi, M.; Ishiyama, K.; Narusaka, Y.; Narusaka, M.; Zhu, J.-K.; Shinozaki, K. Comparative Genomics in Salt Tolerance between Arabidopsis and Arabidopsis-Related Halophyte Salt Cress Using Arabidopsis Microarray. Plant Phys. 2004, 135, 1697–1709. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Glenn, E.P.; O’Leary, J.W. Productivity and irrigation requirements of halophytes grown with seawater in the Sonoran Desert. J. Arid Environ. 1985, 9, 81–91. [Google Scholar] [CrossRef]
- O’Leary, J.W.; Glenn, E.P.; Watson, M.C. Agricultural production of halophytes irrigated with seawater. Plant Soil 1985, 89, 311–321. [Google Scholar]
- Glenn, E.P.; O’Leary, J.W.; Watson, M.C.; Thompson, T.L.; Kuehl, R.O. Salicornia bigelovii Torr.: An Oilseed Halophyte for Seawater Irrigation. Science 1991, 251, 1065–1067. [Google Scholar] [CrossRef] [PubMed]
- Bressan, R.A.; Reddy, M.P.; Chung, S.H.; Yun, D.J.; Hardin, L.S.; Bohnert, H.J. Stress-adapted extremophiles provide energy without interference with food production. Food Sec. 2011, 3, 93–105. [Google Scholar] [CrossRef]
- Das, A.B. Bioprospecting and Genetic Engineering of Mangrove Genes to Enhance Salinity Tolerance in Crop Plants. In Biotechnology of Neglected and Underutilized Crops; Jain, S., Dutta Gupta, S., Eds.; Springer: Dordrecht, The Netherlands, 2013; pp. 385–456. [Google Scholar]
- Rao, N.K.; McCann, I.; Shahid, S.A.; Butt, K.U.R.; Araj, B.A.; Ismail, S. Sustainable use of salt-degraded and abandoned farms for forage production using halophytic grasses. Crop Pasture Sci. 2017, 68, 483. [Google Scholar] [CrossRef]
- Lyra, D.A.; Ismail, S.; Butt, K.U.R.B.; Jed Brown, J. Evaluating the growth performance of eleven Salicornia bigelovii populations under full strength seawater irrigation using multivariate analyses. Aust. J. Crop Sci. 2016, 10, 1429–1441. [Google Scholar] [CrossRef]
- Soni, M.L.; Yadava, N.D.; Kumar, S.; Roy, M.M. Evaluation for growth and yield performance of prickly pear cactus (Opuntia ficus-indica (L.) Mill) accessions in hot arid region of Bikaner, India. Range Manag. Agrofor. 2015, 36, 19–25. [Google Scholar]
- Jellen, E.N.; Maughan, P.J.; Bertero, D.; Munir, H. Prospects for quinoa (Chenopodium Quinoa Willd.) improvement through biotechnology. Biotechnol. Negl. Underutil. Crops 2013, 173–201. [Google Scholar] [CrossRef]
- Yamada, K.; Lim, J.; Dale, J.M.; Chen, H.; Shinn, P.; Palm, C.J.; Southwick, A.M.; Wu, H.C.; Kim, C.; Nguyen, M.; et al. Empirical Analysis of Transcriptional Activity in the Arabidopsis Genome. Science 2003, 302, 842–846. [Google Scholar] [CrossRef] [PubMed]
- Weems, D.; Miller, N.; Garcia-Hernandez, M.; Huala, E.; Rhee, S.Y. Design, Implementation and Maintenance of a Model Organism Database for Arabidopsis thaliana. Comp. Funct. Genom. 2004, 5, 362–369. [Google Scholar] [CrossRef] [PubMed]
- Jaiswal, P.; Avraham, S.; Ilic, K.; Kellogg, E.A.; McCouch, S.; Pujar, A.; Reiser, L.; Rhee, S.Y.; Sachs, M.M.; Schaeffer, M.; et al. Plant Ontology (PO): A Controlled Vocabulary of Plant Structures and Growth Stages. Comp. Funct. Genom. 2005, 6, 388–397. [Google Scholar] [CrossRef] [PubMed]
| In Nature Setting | ||
| Grow Phenotype | Terminology | Survival Phenotype |
| Grow slow | Tolerant | Avoidance of death—lives |
| Grow fast | Sensitive | Avoids avoidance of death—dies |
| Grow fast | Tolerant | Avoidance of death—lives |
| The phenotype in Nature that we seek among species | ||
| In Agriculture/Experimental Setting | ||
| Grow Phenotype | Terminology | Survival Phenotype |
| Grow slow | Sensitive | Avoids avoidance of death—dies |
| Grow fast | Tolerant | Avoidance of death—lives |
| The phenotype in our experiments and screens that we seek and desire for Agriculture use | ||
| Grow slow | Sensitive | Avoids avoidance of death—dies |
| The phenotype that we actually search for in almost all of our experimental screens | ||
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Maggio, A.; Bressan, R.A.; Zhao, Y.; Park, J.; Yun, D.-J. It’s Hard to Avoid Avoidance: Uncoupling the Evolutionary Connection between Plant Growth, Productivity and Stress “Tolerance”. Int. J. Mol. Sci. 2018, 19, 3671. https://doi.org/10.3390/ijms19113671
Maggio A, Bressan RA, Zhao Y, Park J, Yun D-J. It’s Hard to Avoid Avoidance: Uncoupling the Evolutionary Connection between Plant Growth, Productivity and Stress “Tolerance”. International Journal of Molecular Sciences. 2018; 19(11):3671. https://doi.org/10.3390/ijms19113671
Chicago/Turabian StyleMaggio, Albino, Ray A. Bressan, Yang Zhao, Junghoon Park, and Dae-Jin Yun. 2018. "It’s Hard to Avoid Avoidance: Uncoupling the Evolutionary Connection between Plant Growth, Productivity and Stress “Tolerance”" International Journal of Molecular Sciences 19, no. 11: 3671. https://doi.org/10.3390/ijms19113671