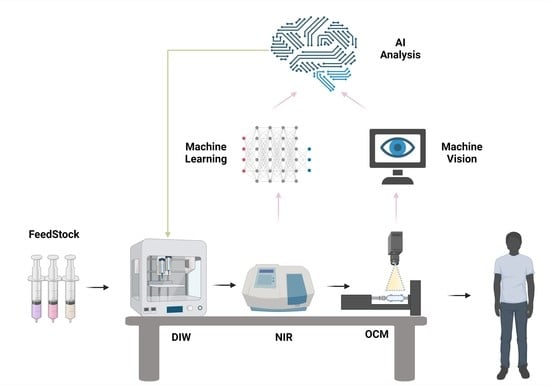
Machine Learning and Machine Vision Accelerate 3D Printed Orodispersible Film Development
Abstract
:1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Methods
2.2.1. Feedstock Formulation
2.2.2. 3D Printing Process
2.2.3. Rheology
2.2.4. Oral Cavity Model (OCM) Disintegration Testing
2.2.5. Petri Dish Method Disintegration
2.2.6. Near-Infrared Spectroscopy Analysis
2.2.7. High-Performance Liquid Chromatography Analysis
2.2.8. Machine Learning
3. Results and Discussion
3.1. 3D Printing Method Development
3.2. ODF Disintegration Study
3.3. Drug and Dose Verification Using ML and NIR
3.4. Classification of ODFs by API
3.5. Regression for Quantifying API Dose
3.6. Realising the Benefits of Digital Technologies
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Liu, J.; Guan, J.; Wan, X.; Shang, R.; Shi, X.; Fang, L.; Liu, C. The Improved Cargo Loading and Physical Stability of Ibuprofen Orodispersible Film: Molecular Mechanism of Ion-Pair Complexes on Drug-Polymer Miscibility. J. Pharm. Sci. 2019, 109, 1356–1364. [Google Scholar] [CrossRef] [PubMed]
- Boulaiz, H.; Ramos, M.C.; Grinan-Lison, C.; Garcia-Rubino, M.E.; Vicente, F.; Marchal, J.A. What’s new in the diagnosis of pancreatic cancer: A patent review (2011-present). Expert Opin. Ther. Pat. 2017, 27, 1319–1328. [Google Scholar] [CrossRef]
- Wang, B.; Yang, L.; Luo, C.; Wang, Y.; Wang, H.; Chen, F.; Xiang, X. Development, In Vitro and In Vivo Evaluation of Racecadotril Orodispersible Films for Pediatric Use. AAPS PharmSciTech 2021, 22, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Gupta, M.S.; Kumar, T.P.; Davidson, R.; Kuppu, G.R.; Pathak, K.; Gowda, D.V. Printing Methods in the Production of Orodispersible Films. AAPS PharmSciTech 2021, 22, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Seoane-Viaño, I.; Trenfield, S.J.; Basit, A.W.; Goyanes, A. Translating 3D printed pharmaceuticals: From hype to real-world clinical applications. Adv. Drug Deliv. Rev. 2021, 174, 553–575. [Google Scholar] [CrossRef] [PubMed]
- Trenfield, S.J.; Xian Tan, H.; Awad, A.; Buanz, A.; Gaisford, S.; Basit, A.W.; Goyanes, A. Track-and-trace: Novel anti-counterfeit measures for 3D printed personalized drug products using smart material inks. Int. J. Pharm. 2019, 567, 118443. [Google Scholar] [CrossRef] [PubMed]
- Seoane-Viaño, I.; Gómez-Lado, N.; Lázare-Iglesias, H.; García-Otero, X.; Antúnez-López, J.R.; Ruibal, Á.; Varela-Correa, J.J.; Aguiar, P.; Basit, A.W.; Otero-Espinar, F.J.; et al. 3D Printed Tacrolimus Rectal Formulations Ameliorate Colitis in an Experimental Animal Model of Inflammatory Bowel Disease. Biomedicines 2020, 8, 563. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; Pose-Boirazian, T.; Eibes, G.; McCoubrey, L.E.; Martínez-Costas, J.; Gaisford, S.; Goyanes, A.; Basit, A.W. A customizable 3D printed device for enzymatic removal of drugs in water. Water Res. 2021, 208, 117861. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; Seijo-Rabina, A.; Awad, A.; Rial, C.; Gaisford, S.; Basit, A.W.; Goyanes, A. Smartphone-enabled 3D printing of medicines. Int. J. Pharm. 2021, 609, 121199. [Google Scholar] [CrossRef] [PubMed]
- Domínguez-Robles, J.; Diaz-Gomez, L.; Utomo, E.; Shen, T.; Picco, C.J.; Alvarez-Lorenzo, C.; Concheiro, A.; Donnelly, R.F.; Larrañeta, E. Use of 3D Printing for the Development of Biodegradable Antiplatelet Materials for Cardiovascular Applications. Pharmaceuticals 2021, 14, 921. [Google Scholar] [CrossRef]
- Falahati, M.; Ahmadvand, P.; Safaee, S.; Chang, Y.-C.; Lyu, Z.; Chen, R.; Li, L.; Lin, Y. Smart polymers and nanocomposites for 3D and 4D printing. Mater. Today 2020, 40, 215–245. [Google Scholar] [CrossRef]
- Jamróz, W.; Kurek, M.; Lyszczarz, E.; Szafraniec-Szczęsny, J.; Knapik-Kowalczuk, J.; Syrek, K.; Paluch, M.; Jachowicz, R. 3D printed orodispersible films with Aripiprazole. Int. J. Pharm. 2017, 533, 413–420. [Google Scholar] [CrossRef]
- Sjöholm, E.; Sandler, N. Additive manufacturing of personalized orodispersible warfarin films. Int. J. Pharm. 2019, 564, 117–123. [Google Scholar] [CrossRef]
- Kiefer, O.; Fischer, B.; Breitkreutz, J. Fundamental Investigations into Metoprolol Tartrate Deposition on Orodispersible Films by Inkjet Printing for Individualised Drug Dosing. Pharmaceutics 2021, 13, 247. [Google Scholar] [CrossRef]
- Thabet, Y.; Lunter, D.; Breitkreutz, J. Continuous inkjet printing of enalapril maleate onto orodispersible film formulations. Int. J. Pharm. 2018, 546, 180–187. [Google Scholar] [CrossRef] [PubMed]
- Edinger, M.; Bar-Shalom, D.; Sandler, N.; Rantanen, J.; Genina, N. QR encoded smart oral dosage forms by inkjet printing. Int. J. Pharm. 2018, 536, 138–145. [Google Scholar] [CrossRef] [PubMed]
- Cho, H.-W.; Baek, S.-H.; Lee, B.-J.; Jin, H.-E. Orodispersible Polymer Films with the Poorly Water-Soluble Drug, Olanzapine: Hot-Melt Pneumatic Extrusion for Single-Process 3D Printing. Pharmaceutics 2020, 12, 692. [Google Scholar] [CrossRef]
- Panraksa, P.; Qi, S.; Udomsom, S.; Tipduangta, P.; Rachtanapun, P.; Jantanasakulwong, K.; Jantrawut, P. Characterization of Hydrophilic Polymers as a Syringe Extrusion 3D Printing Material for Orodispersible Film. Polymers 2021, 13, 3454. [Google Scholar] [CrossRef]
- Karavasili, C.; Eleftheriadis, G.K.; Gioumouxouzis, C.; Andriotis, E.G.; Fatouros, D.G. Mucosal drug delivery and 3D printing technologies: A focus on special patient populations. Adv. Drug Deliv. Rev. 2021, 176, 113858. [Google Scholar] [CrossRef]
- Rahman, J.; Quodbach, J. Versatility on demand—The case for semi-solid micro-extrusion in pharmaceutics. Adv. Drug Deliv. Rev. 2021, 172, 104–126. [Google Scholar] [CrossRef] [PubMed]
- Economidou, S.N.; Uddin, J.; Marques, M.J.; Douroumis, D.; Sow, W.T.; Li, H.; Reid, A.; Windmill, J.F.; Podoleanu, A. A novel 3D printed hollow microneedle microelectromechanical system for controlled, personalized transdermal drug delivery. Addit. Manuf. 2020, 38, 101815. [Google Scholar] [CrossRef]
- Nazir, A.; Azhar, A.; Nazir, U.; Liu, Y.-F.; Qureshi, W.S.; Chen, J.-E.; Alanazi, E. The rise of 3D Printing entangled with smart computer aided design during COVID-19 era. J. Manuf. Syst. 2021, 60, 774–786. [Google Scholar] [CrossRef] [PubMed]
- Beer, N.; Hegger, I.; Kaae, S.; De Bruin, M.L.; Genina, N.; Alves, T.L.; Hoebert, J.; Sporrong, S.K. Scenarios for 3D printing of personalized medicines—A case study. Explor. Res. Clin. Soc. Pharm. 2021, 4, 100073. [Google Scholar] [CrossRef]
- European Medicines Agency. Guideline on Real Time Release Testing (formerly Guideline on Parametric Release). Available online: https://www.ema.europa.eu/en/real-time-release-testing (accessed on 13 December 2021).
- Trenfield, S.J.; Goyanes, A.; Telford, R.; Wilsdon, D.; Rowland, M.; Gaisford, S.; Basit, A.W. 3D printed drug products: Non-destructive dose verification using a rapid point-and-shoot approach. Int. J. Pharm. 2018, 549, 283–292. [Google Scholar] [CrossRef]
- Melocchi, A.; Briatico-Vangosa, F.; Uboldi, M.; Parietti, F.; Turchi, M.; von Zeppelin, D.; Maroni, A.; Zema, L.; Gazzaniga, A.; Zidan, A. Quality considerations on the pharmaceutical applications of fused deposition modeling 3D printing. Int. J. Pharm. 2020, 592, 119901. [Google Scholar] [CrossRef] [PubMed]
- Trenfield, S.J.; Tan, H.X.; Goyanes, A.; Wilsdon, D.; Rowland, M.; Gaisford, S.; Basit, A.W. Non-destructive dose verification of two drugs within 3D printed polyprintlets. Int. J. Pharm. 2020, 577, 119066. [Google Scholar] [CrossRef] [PubMed]
- McCoubrey, L.E.; Gaisford, S.; Orlu, M.; Basit, A.W. Predicting drug-microbiome interactions with machine learning. Biotechnol. Adv. 2021, 107797. [Google Scholar] [CrossRef]
- Elbadawi, M.; McCoubrey, L.E.; Gavins, F.K.; Ong, J.J.; Goyanes, A.; Gaisford, S.; Basit, A.W. Disrupting 3D printing of medicines with machine learning. Trends Pharmacol. Sci. 2021, 42, 745–757. [Google Scholar] [CrossRef]
- McCoubrey, L.; Elbadawi, M.; Orlu, M.; Gaisford, S.; Basit, A. Machine Learning Uncovers Adverse Drug Effects on Intestinal Bacteria. Pharmaceutics 2021, 13, 1026. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Elbadawi, M.; Tsilova, S.L.; Gaisford, S.; Basit, A.W.; Parhizkar, M. Machine learning to empower electrohydrodynamic processing. Mater. Sci. Eng. C 2021, 132, 112553. [Google Scholar] [CrossRef]
- Baker, D.J. Artificial Intelligence: The Future Landscape of Genomic Medical Diagnosis: Dataset, In Silico Artificial Intelligent Clinical Information, and Machine Learning Systems. In Human Genome Informatics. Translating Genes Into Health; Lambert, C.G., Baker, D.J., Patrinos, G.P., Eds.; Elsevier Inc.: Amsterdam, The Netherlands, 2018; pp. 223–267. [Google Scholar]
- Xianyu, Y.; Wang, Q.; Chen, Y. Magnetic particles-enabled biosensors for point-of-care testing. TrAC Trends Anal. Chem. 2018, 106, 213–224. [Google Scholar] [CrossRef]
- Popova, M.; Isayev, O.; Tropsha, A. Deep reinforcement learning for de novo drug design. Sci. Adv. 2018, 4, eaap7885. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hosny, A.; Parmar, C.; Quackenbush, J.; Schwartz, L.H.; Aerts, H.J.W.L. Artificial intelligence in radiology. Nat. Rev. Canc. 2018, 18, 500–510. [Google Scholar] [CrossRef] [PubMed]
- Han, R.; Xiong, H.; Ye, Z.; Yang, Y.; Huang, T.; Jing, Q.; Lu, J.; Pan, H.; Ren, F.; Ouyang, D. Predicting physical stability of solid dispersions by machine learning techniques. J. Control. Release 2019, 311–312, 16–25. [Google Scholar] [CrossRef] [PubMed]
- Harrer, S.; Shah, P.; Antony, B.; Hu, J. Artificial Intelligence for Clinical Trial Design. Trends Pharmacol. Sci. 2019, 40, 577–591. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Elbadawi, M.; Castro, B.M.; Gavins, F.K.; Ong, J.J.; Gaisford, S.; Pérez, G.; Basit, A.W.; Cabalar, P.; Goyanes, A. M3DISEEN: A novel machine learning approach for predicting the 3D printability of medicines. Int. J. Pharm. 2020, 590, 119837. [Google Scholar] [CrossRef] [PubMed]
- Castro, B.M.; Elbadawi, M.; Ong, J.J.; Pollard, T.; Song, Z.; Gaisford, S.; Pérez, G.; Basit, A.W.; Cabalar, P.; Goyanes, A. Machine learning predicts 3D printing performance of over 900 drug delivery systems. J. Control. Release 2021, 337, 530–545. [Google Scholar] [CrossRef] [PubMed]
- Redfearn, A.; Hanson, B. A Mechanical Simulator of Tongue–Palate Compression to Investigate the Oral Flow of Non-Newtonian Fluids. IEEE/ASME Trans. Mechatron. 2018, 23, 958–965. [Google Scholar] [CrossRef]
- Desai, N.; Redfearn, A.; MacLeod, G.; Tuleu, C.; Hanson, B.; Orlu, M. How Do Orodispersible Tablets Behave in an In Vitro Oral Cavity Model: A Pilot Study. Pharmaceutics 2020, 12, 651. [Google Scholar] [CrossRef] [PubMed]
- Levy, G. Viscosity-Stability of Aqueous Solutions of Certain Hydrophilic Polymers. J. Pharm. Sci. 1961, 50, 429–435. [Google Scholar] [CrossRef] [PubMed]
- Benchabane, A.; Bekkour, K. Rheological properties of carboxymethyl cellulose (CMC) solutions. Colloid Polym. Sci. 2008, 286, 1173–1180. [Google Scholar] [CrossRef]
- Elbadawi, M. Polymeric Additive Manufacturing: The Necessity and Utility of Rheology. In Polymer Rheology; Rivera-Armenta, L., Salazar Cruz, B.A., Eds.; IntechOpen: London, UK, 2018; pp. 43–65. [Google Scholar]
- European Pharmacopoeia Commision. European Pharmacopoeia 10.0 Oromucosal Preparations; European Directorate for the Quality of Medicines & Healthcare (EDQM): Strasbourg, France, 2020; Volume 939, pp. 323–324. [Google Scholar]
- Ehtezazi, T.; Algellay, M.; Islam, Y.; Roberts, M.; Dempster, N.M.; Sarker, S.D. The Application of 3D Printing in the Formulation of Multilayered Fast Dissolving Oral Films. J. Pharm. Sci. 2018, 107, 1076–1085. [Google Scholar] [CrossRef] [PubMed]
- Cichocki, A. Unsupervised Learning Algorithms and Latent Variable Models: PCA/SVD, CCA/PLS, ICA, NMF, etc. In Academic Press Library in Signal Processing; Diniz, P.S.R., Suykens, J.A.K., Chellappa, R., Theodoridis, S., Eds.; Elsevier Inc.: Amsterdam, The Netherlands, 2014; Volume 1, pp. 1151–1238. [Google Scholar]
- Moncayo, S.; Manzoor, S.; Navarro-Villoslada, F.; Caceres, J. Evaluation of supervised chemometric methods for sample classification by Laser Induced Breakdown Spectroscopy. Chemom. Intell. Lab. Syst. 2015, 146, 354–364. [Google Scholar] [CrossRef]
- Clark, M.; Cramer, R.D. The Probability of Chance Correlation Using Partial Least Squares (PLS). Quant. Struct. Relationships 1993, 12, 137–145. [Google Scholar] [CrossRef]
- Elbadawi, M.; McCoubrey, L.E.; Gavins, F.K.; Ong, J.J.; Goyanes, A.; Gaisford, S.; Basit, A.W. Harnessing artificial intelligence for the next generation of 3D printed medicines. Adv. Drug Deliv. Rev. 2021, 175, 113805. [Google Scholar] [CrossRef] [PubMed]
- Raijada, D.; Wac, K.; Greisen, E.; Rantanen, J.; Genina, N. Integration of personalized drug delivery systems into digital health. Adv. Drug Deliv. Rev. 2021, 176, 113857. [Google Scholar] [CrossRef]
- Awad, A.; Trenfield, S.J.; Pollard, T.D.; Ong, J.J.; Elbadawi, M.; McCoubrey, L.E.; Goyanes, A.; Gaisford, S.; Basit, A.W. Connected healthcare: Improving patient care using digital health technologies. Adv. Drug Deliv. Rev. 2021, 178, 113958. [Google Scholar] [CrossRef] [PubMed]












| Feedstock Formulation | Water (mL) | API (g) | CMC (g) |
|---|---|---|---|
| 5% w/w Paracetamol | 100 | 0.13 | 2.5 |
| 10% w/w Paracetamol | 100 | 0.28 | 2.5 |
| 20% w/w Paracetamol | 100 | 0.625 | 2.5 |
| 5% w/w Caffeine | 100 | 0.13 | 2.5 |
| 10% w/w Caffeine | 100 | 0.28 | 2.5 |
| 20% w/w Caffeine | 100 | 0.625 | 2.5 |
| 5% w/w Theophylline | 100 | 0.13 | 2.5 |
| 10% w/w Theophylline | 100 | 0.28 | 2.5 |
| 20% w/w Theophylline | 100 | 0.625 | 2.5 |
| Parameters | |
|---|---|
| Needle gauge (Needle diameter) | 22G (0.410 mm) |
| Compressed air pressure | 100 kPa |
| Printing speed | 20 mm/s |
| Infill pattern | Grid infill |
| Infill density | 25% |
| Parameters | Paracetamol | Caffeine | Theophylline |
|---|---|---|---|
| Mobile phase composition | A: Distilled Water | A: Orthophosphoric Acid | A: Distilled Water |
| B: Methanol | B: Acetonitrile | B: Acetonitrile | |
| C: Ethanol | |||
| Mobile Phase ratio | 85:15 | 80:20 | 60:10:30 |
| Flow rate (mL/min) | 1 | 1 | 1 |
| Injection volume (µL) | 20 | 10 | 1 |
| Detection wavelength (nm) | 247 | 272 | 272 |
| Column | Luna C18 (250 × 4.6 mm; 5 µm) | Luna C18 (250 × 4.6 mm; 5 µm) | Luna C18 (250 × 4.6 mm; 5 µm) |
| Column temperature (°C) | 40 | 40 | 40 |
| Retention time (mins) | 8.8 | 13.0 | 13.3 |
| Strip (25% ID) | Strip (40% ID) | Star (25% ID) | Circle (25% ID) | |
|---|---|---|---|---|
| 0 s |  |  |  |  |
| 60 s |  |  |  |  |
| 120 s |  |  |  |  |
| 180 s |  |  |  |  |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
O’Reilly, C.S.; Elbadawi, M.; Desai, N.; Gaisford, S.; Basit, A.W.; Orlu, M. Machine Learning and Machine Vision Accelerate 3D Printed Orodispersible Film Development. Pharmaceutics 2021, 13, 2187. https://doi.org/10.3390/pharmaceutics13122187
O’Reilly CS, Elbadawi M, Desai N, Gaisford S, Basit AW, Orlu M. Machine Learning and Machine Vision Accelerate 3D Printed Orodispersible Film Development. Pharmaceutics. 2021; 13(12):2187. https://doi.org/10.3390/pharmaceutics13122187
Chicago/Turabian StyleO’Reilly, Colm S., Moe Elbadawi, Neel Desai, Simon Gaisford, Abdul W. Basit, and Mine Orlu. 2021. "Machine Learning and Machine Vision Accelerate 3D Printed Orodispersible Film Development" Pharmaceutics 13, no. 12: 2187. https://doi.org/10.3390/pharmaceutics13122187