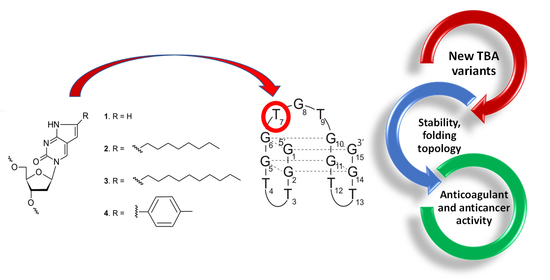
A Comprehensive Analysis of the Thrombin Binding Aptamer Containing Functionalized Pyrrolo-2’-deoxycytidines
Abstract
:1. Introduction
2. Results and Discussion
2.1. Thermodynamic Properties of TBA Variants
2.2. The Analysis of TBA Variants Folding Topology
2.3. Anticoagulant Properties of Modified Thrombin Binding Aptamer Variants
2.4. Antiproliferative Properties of Modified Thrombin Binding Aptamer Variants
3. Materials and Methods
3.1. Chemical Synthesis of Oligonucleotides
3.2. UV Melting Analysis
3.3. Circular Dichroism Spectra
3.4. Thermal Difference Spectra
3.5. Thrombin Time Assay
3.6. The Real-Time Cellular Impedance Assay
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Santosh, B.; Yadava, P.K. Nucleic Acid Aptamers: Research Tools in Disease Diagnostics and Therapeutics. BioMed Res. Int. 2014, 540451, 1–13. [Google Scholar] [CrossRef]
- Kotkowiak, W.; Pasternak, A. Beyond G-Quadruplexes—The Effect of Junction with Additional Structural Motifs on Aptamers Properties. Int. J. Mol. Sci. 2021, 22, 9948. [Google Scholar] [CrossRef] [PubMed]
- Zhang, N.; Chen, Z.; Liu, D.; Jiang, H.; Zhang, Z.-K.; Lu, A.; Zhang, B.-T.; Yu, Y.; Zhang, G. Structural Biology for the Molecular Insight between Aptamers and Target Proteins. Int. J. Mol. Sci. 2021, 22, 4093. [Google Scholar] [CrossRef]
- Gelinas, A.D.; Davies, D.R.; Janjic, N. Embracing Proteins: Structural Themes in Aptamer-Protein Complexes. Curr. Opin. Struct. Biol. 2016, 36, 122–132. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bock, L.C.; Griffin, L.C.; Latham, J.A.; Vermaas, E.H.; Toole, J.J. Selection of Single-Stranded DNA Molecules That Bind and Inhibit Human Thrombin. Nature 1992, 355, 564–566. [Google Scholar] [CrossRef]
- Wang, K.Y.; Krawczyk, S.H.; Bischofberger, N.; Swaminathan, S.; Bolton, P.H. The Tertiary Structure of a DNA Aptamer Which Binds to and Inhibits Thrombin Determines Activity. Biochemistry 1993, 32, 11285–11292. [Google Scholar] [CrossRef]
- Padmanabhan, K.; Tulinsky, A. An Ambiguous Structure of a DNA 15-Mer Thrombin Complex. Acta Crystallogr. 1996, 52, 272–282. [Google Scholar] [CrossRef] [Green Version]
- Posma, J.J.N.; Posthuma, J.J.; Spronk, H.M.H. Coagulation and Non-Coagulation Effects of Thrombin. J. Thromb. Haemost. 2016, 14, 1908–1916. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Adams, G.N.; Rosenfeldt, L.; Frederick, M.; Miller, W.; Waltz, D.; Kombrinck, K.; McElhinney, K.E.; Flick, M.J.; Monia, B.P.; Revenko, A.S.; et al. Colon Cancer Growth and Dissemination Relies upon Thrombin, Stromal PAR-1, and Fibrinogen. Cancer Res. 2015, 75, 4235–4243. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, S.; Xiao, W.; Pan, X.; Zhu, M.; Yang, Z.; Zhang, F.; Zheng, C. Thrombin Promotes Proliferation of Human Lung Fibroblasts via Protease Activated Receptor-1-Dependent and NF-ΚB-Independent Pathways. Cell Biol. Int. 2014, 38, 747–756. [Google Scholar] [CrossRef]
- Coppens, M.; Eikelboom, J.W.; Gustafsson, D.; Weitz, J.I.; Hirsh, J. Translational Success Stories: Development of Direct Thrombin Inhibitors. Circ. Res. 2012, 111, 920–929. [Google Scholar] [CrossRef] [Green Version]
- Crawley, J.T.B.; Zanardelli, S.; Chion, C.K.N.K.; Lane, D.A. The Central Role of Thrombin in Hemostasis. J. Thromb. Haemost. 2007, 5, 95–101. [Google Scholar] [CrossRef]
- Russo Krauss, I.; Merlino, A.; Giancola, C.; Randazzo, A.; Mazzarella, L.; Sica, F. Thrombin-Aptamer Recognition: A Revealed Ambiguity. Nucleic Acids Res. 2011, 39, 7858–7867. [Google Scholar] [CrossRef] [Green Version]
- Pica, A.; Russo Krauss, I.; Merlino, A.; Nagatoishi, S.; Sugimoto, N.; Sica, F. Dissecting the Contribution of Thrombin Exosite I in the Recognition of Thrombin Binding Aptamer. FEBS J. 2013, 280, 6581–6588. [Google Scholar] [CrossRef]
- Mendelboum Raviv, S.; Horváth, A.; Aradi, J.; Bagoly, Z.; Fazakas, F.; Batta, Z.; Muszbek, L.; Hársfalvi, J. 4-Thio-Deoxyuridylate-Modified Thrombin Aptamer and Its Inhibitory Effect on Fibrin Clot Formation, Platelet Aggregation and Thrombus Growth on Subendothelial Matrix. J. Thromb. Haemost. 2008, 6, 1764–1771. [Google Scholar] [CrossRef] [PubMed]
- Kotkowiak, W.; Czapik, T.; Pasternak, A. Novel Isoguanine Derivative of Unlocked Nucleic Acid-Investigations of Thermodynamics and Biological Potential of Modified Thrombin Binding Aptamer. PLoS ONE 2018, 13, 0197835–0197849. [Google Scholar] [CrossRef]
- Nallagatla, S.R.; Heuberger, B.; Haque, A.; Switzer, C. Combinatorial Synthesis of Thrombin-Binding Aptamers Containing Iso-Guanine. J. Comb. Chem. 2009, 11, 364–369. [Google Scholar] [CrossRef] [PubMed]
- Bonifacio, L.; Church, F.C.; Jarstfer, M.B. Effect of Locked-Nucleic Acid on a Biologically Active g-Quadruplex. A Structure-Activity Relationship of the Thrombin Aptamer. Int. J. Mol. Sci. 2008, 9, 422–433. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zaitseva, M.; Kaluzhny, D.; Shchyolkina, A.; Borisova, O.; Smirnov, I.; Pozmogova, G. Conformation and Thermostability of Oligonucleotide d(GGTTGGTGTGGTTGG) Containing Thiophosphoryl Internucleotide Bonds at Different Positions. Biophys. Chem. 2010, 146, 1–6. [Google Scholar] [CrossRef]
- Smirnov, I.; Shafer, R.H. Effect of Loop Sequence and Size on DNA Aptamer Stability. Biochemistry 2000, 39, 1462–1468. [Google Scholar] [CrossRef] [PubMed]
- Scuotto, M.; Rivieccio, E.; Varone, A.; Corda, D.; Bucci, M.; Vellecco, V.; Cirino, G.; Virgilio, A.; Esposito, V.; Galeone, A.; et al. Site Specific Replacements of a Single Loop Nucleoside with a Dibenzyl Linker May Switch the Activity of TBA from Anticoagulant to Antiproliferative. Nucleic Acids Res. 2015, 43, 7702–7716. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pecoraro, A.; Virgilio, A.; Esposito, V.; Galeone, A.; Russo, G.; Russo, A. UL3 Mediated Nucleolar Stress Pathway as a New Mechanism of Action of Antiproliferative G-Quadruplex TBA Derivatives in Colon Cancer Cells. Biomolecules 2020, 10, 583. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Esposito, V.; Russo, A.; Vellecco, V.; Bucci, M.; Russo, G.; Mayol, L.; Virgilio, A.; Galeone, A. Thrombin Binding Aptamer Analogues Containing Inversion of Polarity Sites Endowed with Antiproliferative and Anti-Motility Properties against Calu-6 Cells. Biochim. Biophys. Acta. Gen. Subj. 2018, 1862, 2645–2650. [Google Scholar] [CrossRef]
- Esposito, V.; Russo, A.; Amato, T.; Vellecco, V.; Bucci, M.; Mayol, L.; Russo, G.; Virgilio, A.; Galeone, A. The “Janus Face” of the Thrombin Binding Aptamer: Investigating the Anticoagulant and Antiproliferative Properties through Straightforward Chemical Modifications. Bioorg. Chem. 2018, 76, 202–209. [Google Scholar] [CrossRef] [PubMed]
- Virgilio, A.; Esposito, V.; Pecoraro, A.; Russo, A.; Vellecco, V.; Pepe, A.; Bucci, M.; Russo, G.; Galeone, A. Structural Properties and Anticoagulant/Cytotoxic Activities of Heterochiral Enantiomeric Thrombin Binding Aptamer (TBA) Derivatives. Nucleic Acids Res. 2020, 48, 12556–12565. [Google Scholar] [CrossRef] [PubMed]
- Riccardi, C.; Meyer, A.; Vasseur, J.-J.; Russo Krauss, I.; Paduano, L.; Morvan, F.; Montesarchio, D. Fine-Tuning the Properties of the Thrombin Binding Aptamer through Cyclization: Effect of the 5’-3’ Connecting Linker on the Aptamer Stability and Anticoagulant Activity. Bioorg. Chem. 2020, 94, 103379. [Google Scholar] [CrossRef] [PubMed]
- Bao, H.-L.; Ishizuka, T.; Yamashita, A.; Furukoji, E.; Asada, Y.; Xu, Y. Improving Thermodynamic Stability and Anticoagulant Activity of a Thrombin Binding Aptamer by Incorporation of 8-Trifluoromethyl-2’-Deoxyguanosine. J. Med. Chem. 2021, 64, 711–718. [Google Scholar] [CrossRef] [PubMed]
- Berry, D.A.; Jung, K.-Y.; Wise, D.S.; Sercel, A.D.; Pearson, W.H.; Mackie, H.; Randolph, J.B.; Somers, R.L. Pyrrolo-DC and Pyrrolo-C: Fluorescent Analogs of Cytidine and 2′-Deoxycytidine for the Study of Oligonucleotides. Tetrahedron Lett. 2004, 45, 2457–2461. [Google Scholar] [CrossRef]
- Jahnz-Wechmann, Z.; Lisowiec-Wachnicka, J.; Framski, G.; Kosman, J.; Boryski, J.; Pasternak, A. Thermodynamic, Structural and Fluorescent Characteristics of DNA Hairpins Containing Functionalized Pyrrolo-2’-Deoxycytidines. Bioorg. Chem. 2017, 71, 294–298. [Google Scholar] [CrossRef]
- Pasternak, A.; Hernandez, F.J.; Rasmussen, L.M.; Vester, B.; Wengel, J. Improved Thrombin Binding Aptamer by Incorporation of a Single Unlocked Nucleic Acid Monomer. Nucleic Acids Res. 2011, 39, 1155–1164. [Google Scholar] [CrossRef] [Green Version]
- Kotkowiak, W.; Lisowiec-Wachnicka, J.; Grynda, J.; Kierzek, R.; Wengel, J.; Pasternak, A. Thermodynamic, Anticoagulant, and Antiproliferative Properties of Thrombin Binding Aptamer Containing Novel UNA Derivative. Mol. Ther. Nucleic Acids 2018, 10, 304–316. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Borbone, N.; Bucci, M.; Oliviero, G.; Morelli, E.; Amato, J.; D’Atri, V.; D’Errico, S.; Vellecco, V.; Cirino, G.; Piccialli, G.; et al. Investigating the Role of T7 and T12 Residues on the Biological Properties of Thrombin-Binding Aptamer: Enhancement of Anticoagulant Activity by a Single Nucleobase Modification. J. Med. Chem. 2012, 55, 10716–10728. [Google Scholar] [CrossRef]
- Migliore, M.D.; Zonta, N.; McGuigan, C.; Henson, G.; Andrei, G.; Snoeck, R.; Balzarini, J. Synthesis and Antiviral Activity of the Carbocyclic Analogue of the Highly Potent and Selective Anti-VZV Bicyclo Furano Pyrimidines. J. Med. Chem. 2007, 50, 6485–6492. [Google Scholar] [CrossRef] [PubMed]
- Russo Krauss, I.; Merlino, A.; Randazzo, A.; Novellino, E.; Mazzarella, L.; Sica, F. High-Resolution Structures of Two Complexes between Thrombin and Thrombin-Binding Aptamer Shed Light on the Role of Cations in the Aptamer Inhibitory Activity. Nucleic Acids Res. 2012, 40, 8119–8128. [Google Scholar] [CrossRef] [Green Version]
- Giraldo, R.; Suzuki, M.; Chapman, L.; Rhodes, D. Promotion of Parallel DNA Quadruplexes by a Yeast Telomere Binding Protein: A Circular Dichroism Study. Proc. Natl. Acad. Sci. USA 1994, 91, 7658–7662. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mergny, J.-L.; Li, J.; Lacroix, L.; Amrane, S.; Chaires, J.B. Thermal Difference Spectra: A Specific Signature for Nucleic Acid Structures. Nucleic Acids Res. 2005, 33, 138–144. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kotkowiak, W.; Wengel, J.; Scotton, C.J.; Pasternak, A. Improved RE31 Analogues Containing Modified Nucleic Acid Monomers: Thermodynamic, Structural, and Biological Effects. J. Med. Chem. 2019, 62, 2499–2507. [Google Scholar] [CrossRef] [PubMed]
- Nagatoishi, S.; Isono, N.; Tsumoto, K.; Sugimoto, N. Loop Residues of Thrombin-Binding DNA Aptamer Impact G-Quadruplex Stability and Thrombin Binding. Biochimie 2011, 93, 1231–1238. [Google Scholar] [CrossRef] [PubMed]
- Pagano, B.; Martino, L.; Randazzo, A.; Giancola, C. Stability and Binding Properties of a Modified Thrombin Binding Aptamer. Biophys. J. 2008, 94, 562–569. [Google Scholar] [CrossRef] [Green Version]
- Cai, B.; Yang, X.; Sun, L.; Fan, X.; Li, L.; Jin, H.; Wu, Y.; Guan, Z.; Zhang, L.; Zhang, L.; et al. Stability and Bioactivity of Thrombin Binding Aptamers Modified with D-/L-Isothymidine in the Loop Regions. Org. Biomol. Chem. 2014, 12, 8866–8876. [Google Scholar] [CrossRef] [PubMed]
- Bock, P.E.; Panizzi, P.; Verhamme, I.M.A. Exosites in the Substrate Specificity of Blood Coagulation Reactions. J. Thromb. Haemost. 2007, 5, 81–94. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tsvetkov, V.B.; Varizhuk, A.M.; Pozmogova, G.E.; Smirnov, I.P.; Kolganova, N.A.; Timofeev, E.N. A Universal Base in a Specific Role: Tuning up a Thrombin Aptamer with 5-Nitroindole. Sci. Rep. 2015, 5, 16337–16348. [Google Scholar] [CrossRef]
- Bates, P.J.; Laber, D.A.; Miller, D.M.; Thomas, S.D.; Trent, J.O. Discovery and Development of the G-Rich Oligonucleotide AS1411 as a Novel Treatment for Cancer. Exp. Mol. Pathol. 2009, 86, 151–164. [Google Scholar] [CrossRef] [Green Version]
- Choi, E.W.; Nayak, L.V.; Bates, P.J. Cancer-Selective Antiproliferative Activity Is a General Property of Some G-Rich Oligodeoxynucleotides. Nucleic Acids Res. 2010, 38, 1623–1635. [Google Scholar] [CrossRef]
- Roxo, C.; Kotkowiak, W.; Pasternak, A. G4 Matters—the Influence of G-Quadruplex Structural Elements on the Antiproliferative Properties of G-Rich Oligonucleotides. Int. J. Mol. Sci. 2021, 22, 4941. [Google Scholar] [CrossRef]
- Roy, S.; Caruthers, M. Synthesis of DNA/RNA and Their Analogs via Phosphoramidite and H-Phosphonate Chemistries. Molecules 2013, 18, 14268–14284. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lopez-Gomollon, S.; Nicolas, F.E. Purification of DNA Oligos by Denaturing Polyacrylamide Gel Electrophoresis (PAGE). Meth. Enzymol. 2013, 529, 65–83. [Google Scholar]
- Mergny, J.-L.; Lacroix, L. UV Melting of G-Quadruplexes. Curr. Protoc. Nucleic Acid Chem. 2009, 17, 1–15. [Google Scholar] [CrossRef] [PubMed]



| Name | Sequence (5ʹ-3ʹ) | Average of Curve Fits a | |||||
|---|---|---|---|---|---|---|---|
| −ΔH° (kcal/mol) | −ΔS° (eu) | ΔG°37 (kcal/mol) | TM (°C) | ΔΔG°37 (kcal/mol) | ΔTM (°C) | ||
| TBA | GGTTGGTGTGGTTGG | 41.2 ± 0.9 | 127.2 ± 2.7 | −1.74 ± 0.02 | 50.7 | 0 | 0 |
| ON1 | GGTTGG1GTGGTTGG | 37.2 ± 0.5 | 115.3 ± 1.5 | −1.49 ± 0.03 | 50.0 | 0.25 | −0.7 |
| ON2 | GGTTGG2GTGGTTGG | 38.6 ± 1.5 | 119.5 ± 4.8 | −1.57 ± 0.05 | 50.1 | 0.17 | −0.6 |
| ON3 | GGTTGG3GTGGTTGG | 39.6 ± 1.5 | 122.7 ± 4.7 | −1.51 ± 0.03 | 49.3 | 0.23 | −1.4 |
| ON4 | GGTTGG4GTGGTTGG | 29.6 ± 1.1 | 91.7 ± 3.6 | −1.17 ± 0.02 | 49.8 | 0.57 | −0.9 |
| Oligomer | Sequence | AE a (s) |
|---|---|---|
| TBA | GGTTGGTGTGGTTGG | 26.2 |
| ON1 | GGTTGG1GTGGTTGG | 8.0 |
| ON2 | GGTTGG2GTGGTTGG | 20.0 |
| ON3 | GGTTGG3GTGGTTGG | 30.0 |
| ON4 | GGTTGG4GTGGTTGG | 9.4 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kotkowiak, W.; Jahnz-Wechmann, Z.; Pasternak, A. A Comprehensive Analysis of the Thrombin Binding Aptamer Containing Functionalized Pyrrolo-2’-deoxycytidines. Pharmaceuticals 2021, 14, 1326. https://doi.org/10.3390/ph14121326
Kotkowiak W, Jahnz-Wechmann Z, Pasternak A. A Comprehensive Analysis of the Thrombin Binding Aptamer Containing Functionalized Pyrrolo-2’-deoxycytidines. Pharmaceuticals. 2021; 14(12):1326. https://doi.org/10.3390/ph14121326
Chicago/Turabian StyleKotkowiak, Weronika, Zofia Jahnz-Wechmann, and Anna Pasternak. 2021. "A Comprehensive Analysis of the Thrombin Binding Aptamer Containing Functionalized Pyrrolo-2’-deoxycytidines" Pharmaceuticals 14, no. 12: 1326. https://doi.org/10.3390/ph14121326